[English] 日本語
 Yorodumi
Yorodumi- EMDB-41810: Cryo-EM structure of vaccine-elicited CD4 binding site antibody D... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement | |||||||||
 Map data Map data | Locally refined map showing the vaccine elicited DH1285 antibody fab bound to one of pg120 protomer of CH505M5chimer.6R.SOSIP.664v4.1 Env | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HIV-1 Vaccine induced neutralizing antibody / CD4 binding site antibody / gp120 / DH1285 / IgG / rhesus macaques / CH505M5chimer.6R.SOSIP.664v4.1 Env / VIRAL PROTEIN | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.25 Å | |||||||||
 Authors Authors | Thakur B / Stalls VD / Acharya P | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: Vaccine induction of CD4-mimicking HIV-1 broadly neutralizing antibody precursors in macaques. Authors: Kevin O Saunders / James Counts / Bhishem Thakur / Victoria Stalls / Robert Edwards / Kartik Manne / Xiaozhi Lu / Katayoun Mansouri / Yue Chen / Rob Parks / Maggie Barr / Laura Sutherland / ...Authors: Kevin O Saunders / James Counts / Bhishem Thakur / Victoria Stalls / Robert Edwards / Kartik Manne / Xiaozhi Lu / Katayoun Mansouri / Yue Chen / Rob Parks / Maggie Barr / Laura Sutherland / Joena Bal / Nicholas Havill / Haiyan Chen / Emily Machiele / Nolan Jamieson / Bhavna Hora / Megan Kopp / Katarzyna Janowska / Kara Anasti / Chuancang Jiang / Elizabeth Van Itallie / Sravani Venkatayogi / Amanda Eaton / Rory Henderson / Christopher Barbosa / S Munir Alam / Sampa Santra / Drew Weissman / M Anthony Moody / Derek W Cain / Ying K Tam / Mark Lewis / Wilton B Williams / Kevin Wiehe / David C Montefiori / Priyamvada Acharya / Barton F Haynes /   Abstract: The CD4-binding site (CD4bs) is a conserved epitope on HIV-1 envelope (Env) that can be targeted by protective broadly neutralizing antibodies (bnAbs). HIV-1 vaccines have not elicited CD4bs bnAbs ...The CD4-binding site (CD4bs) is a conserved epitope on HIV-1 envelope (Env) that can be targeted by protective broadly neutralizing antibodies (bnAbs). HIV-1 vaccines have not elicited CD4bs bnAbs for many reasons, including the occlusion of CD4bs by glycans, expansion of appropriate naive B cells with immunogens, and selection of functional antibody mutations. Here, we demonstrate that immunization of macaques with a CD4bs-targeting immunogen elicits neutralizing bnAb precursors with structural and genetic features of CD4-mimicking bnAbs. Structures of the CD4bs nAb bound to HIV-1 Env demonstrated binding angles and heavy-chain interactions characteristic of all known human CD4-mimicking bnAbs. Macaque nAb were derived from variable and joining gene segments orthologous to the genes of human VH1-46-class bnAb. This vaccine study initiated in primates the B cells from which CD4bs bnAbs can derive, accomplishing the key first step in the development of an effective HIV-1 vaccine. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41810.map.gz emd_41810.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41810-v30.xml emd-41810-v30.xml emd-41810.xml emd-41810.xml | 21.7 KB 21.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_41810.png emd_41810.png | 23.2 KB | ||
| Filedesc metadata |  emd-41810.cif.gz emd-41810.cif.gz | 7.1 KB | ||
| Others |  emd_41810_half_map_1.map.gz emd_41810_half_map_1.map.gz emd_41810_half_map_2.map.gz emd_41810_half_map_2.map.gz | 115.5 MB 115.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41810 http://ftp.pdbj.org/pub/emdb/structures/EMD-41810 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41810 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41810 | HTTPS FTP |
-Validation report
| Summary document |  emd_41810_validation.pdf.gz emd_41810_validation.pdf.gz | 883.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41810_full_validation.pdf.gz emd_41810_full_validation.pdf.gz | 883.5 KB | Display | |
| Data in XML |  emd_41810_validation.xml.gz emd_41810_validation.xml.gz | 13.9 KB | Display | |
| Data in CIF |  emd_41810_validation.cif.gz emd_41810_validation.cif.gz | 16.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41810 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41810 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41810 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41810 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41810.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41810.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally refined map showing the vaccine elicited DH1285 antibody fab bound to one of pg120 protomer of CH505M5chimer.6R.SOSIP.664v4.1 Env | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
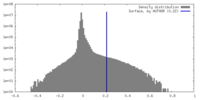
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_41810_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
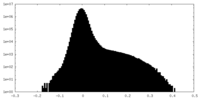
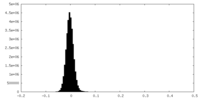
| Density Histograms |
-Half map: #1
| File | emd_41810_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
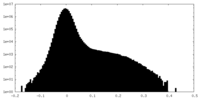
| Density Histograms |
- Sample components
Sample components
-Entire : The complex of vaccine elicited CD4 binding site neutralizing ant...
| Entire | Name: The complex of vaccine elicited CD4 binding site neutralizing antibody DH1285 Fab with CH505M5chimer.6R.SOSIP.664v4.1 Env |
|---|---|
| Components |
|
-Supramolecule #1: The complex of vaccine elicited CD4 binding site neutralizing ant...
| Supramolecule | Name: The complex of vaccine elicited CD4 binding site neutralizing antibody DH1285 Fab with CH505M5chimer.6R.SOSIP.664v4.1 Env type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: One of the gp120 protomer bound to DH1285 locally refined to get precise information about the interacting residues. |
|---|---|
| Molecular weight | Theoretical: 308 KDa |
-Supramolecule #2: Envelope glycoprotein gp120
| Supramolecule | Name: Envelope glycoprotein gp120 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Supramolecule #3: DH1285 Heavy Chain, DH1285 Light Chain
| Supramolecule | Name: DH1285 Heavy Chain, DH1285 Light Chain / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 1 / Details: CH505M5chimer.6R.SOSIP.664v4.1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 72.669078 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPMGSLQPLA TLYLLGMLVA SVLAAENLWV TVYYGVPVWK EAKTTLFCAS DAKAYEKKVH NVWATHACVP TDPNPQEMVL KNVTENFNM WKNDMVDQMH EDVISLWDQS LKPCVKLTPL CVTLNCTNAT ASNSSIIEGM KNCSFNITTE LRDKREKKNA L FYKLDIVQ ...String: MPMGSLQPLA TLYLLGMLVA SVLAAENLWV TVYYGVPVWK EAKTTLFCAS DAKAYEKKVH NVWATHACVP TDPNPQEMVL KNVTENFNM WKNDMVDQMH EDVISLWDQS LKPCVKLTPL CVTLNCTNAT ASNSSIIEGM KNCSFNITTE LRDKREKKNA L FYKLDIVQ LDGNSSQYRL INCNTSVITQ ACPKVSFDPI PIHYCAPAGY AILKCNNKTF TGTGPCNNVS TVQCTHGIKP VV STQLLLN GSLAEGEIII RSENITNNVK TIIVHLNESV KIECTRPNNK TRTSIRIGPG QWFYATGQVI GDIREAYCNI NES KWNETL QRVSKKLKEY FPHKNITFQP SSGGDLEITT HSFNCGGEFF YCNTSSLFNR TYMANSTDMA NSTETNSTRT ITIH CRIKQ IINMWQEVGR AMYAPPIAGN ITCISNITGL LLTRDGGKNN TETFRPGGGN MKDNWRSELY KYKVVKIEPL GVAPT RCKR RVVGRRRRRR AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RNLLSGIVQQ QSNLLRAPEA QQHLLKLTVW GIKQLQ ARV LAVERYLRDQ QLLGIWGCSG KLICCTNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNE QD LLALD |
-Macromolecule #2: DH1285 Heavy Chain
| Macromolecule | Name: DH1285 Heavy Chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.396316 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGWSCIILFL VATATGVHAQ VHLEQSGAEV KEPGSSVRLS CEASGYTFTD YYIHWVRQSP RQGLEWMGWI NPYYGNTHYA EKFQGRVAM TRDRSTTTAY MDLSSLTSED TAVYYCARDE GGSGSYSYFD SWGQGVLVTV SSASTKGPSV FPLAPSSRST S ESTAALGC ...String: MGWSCIILFL VATATGVHAQ VHLEQSGAEV KEPGSSVRLS CEASGYTFTD YYIHWVRQSP RQGLEWMGWI NPYYGNTHYA EKFQGRVAM TRDRSTTTAY MDLSSLTSED TAVYYCARDE GGSGSYSYFD SWGQGVLVTV SSASTKGPSV FPLAPSSRST S ESTAALGC LVKDYFPEPV TVSWNSGSLT SGVHTFPAVL QSSGLYSLSS VVTVPSSSLG TQTYVCNVNH KPSNTKVDKR VE IKT |
-Macromolecule #3: DH1285 Light Chain
| Macromolecule | Name: DH1285 Light Chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.240955 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGWSCIILFL VATATGVHAD IQMTQSPSSL SASVGDTVTI TCRASQDINN HLSWYQQKPG RAPKALIYSA SSLETGVPSR FSGSGSGTD YTLTISSLQP EDFATYYCQH YSTSPYTFGR GTKVDIKRAV AAPSVFIFPP SEDQVKSGTV SVVCLLNNFY P REASVKWK ...String: MGWSCIILFL VATATGVHAD IQMTQSPSSL SASVGDTVTI TCRASQDINN HLSWYQQKPG RAPKALIYSA SSLETGVPSR FSGSGSGTD YTLTISSLQP EDFATYYCQH YSTSPYTFGR GTKVDIKRAV AAPSVFIFPP SEDQVKSGTV SVVCLLNNFY P REASVKWK VDGVLKTGNS QESVTEQDSK DNTYSLSSTL TLSSTDYQSH NVYACEVTHQ GLSSPVTKSF NRGEC |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 295 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 59.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8u1d: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)