[English] 日本語
 Yorodumi
Yorodumi- EMDB-41071: Cryo-EM structure of rat cardiac sodium channel NaV1.5 with batra... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of rat cardiac sodium channel NaV1.5 with batrachotoxin analog BTX-B | |||||||||||||||
 Map data Map data | Sharpened map CryoSPARC | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Ion channel / Sodium channel / Voltage-gated channel / Sodium transport / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcell communication involved in cardiac conduction / voltage-gated sodium channel activity involved in AV node cell action potential / voltage-gated sodium channel activity involved in bundle of His cell action potential / voltage-gated sodium channel activity involved in Purkinje myocyte action potential / voltage-gated sodium channel activity involved in SA node cell action potential / bundle of His cell action potential / regulation of ventricular cardiac muscle cell membrane depolarization / AV node cell action potential / SA node cell action potential / AV node cell to bundle of His cell communication ...cell communication involved in cardiac conduction / voltage-gated sodium channel activity involved in AV node cell action potential / voltage-gated sodium channel activity involved in bundle of His cell action potential / voltage-gated sodium channel activity involved in Purkinje myocyte action potential / voltage-gated sodium channel activity involved in SA node cell action potential / bundle of His cell action potential / regulation of ventricular cardiac muscle cell membrane depolarization / AV node cell action potential / SA node cell action potential / AV node cell to bundle of His cell communication / membrane depolarization during SA node cell action potential / response to denervation involved in regulation of muscle adaptation / membrane depolarization during atrial cardiac muscle cell action potential / sodium channel complex / cardiac ventricle development / voltage-gated sodium channel activity involved in cardiac muscle cell action potential / regulation of atrial cardiac muscle cell membrane repolarization / brainstem development / membrane depolarization during AV node cell action potential / regulation of atrial cardiac muscle cell membrane depolarization / positive regulation of action potential / membrane depolarization during bundle of His cell action potential / atrial cardiac muscle cell action potential / membrane depolarization during Purkinje myocyte cell action potential / telencephalon development / membrane depolarization during cardiac muscle cell action potential / membrane depolarization during action potential / positive regulation of sodium ion transport / regulation of sodium ion transmembrane transport / ventricular cardiac muscle cell action potential / regulation of ventricular cardiac muscle cell membrane repolarization / cardiac muscle cell action potential involved in contraction / voltage-gated sodium channel complex / regulation of cardiac muscle cell contraction / sodium ion import across plasma membrane / voltage-gated sodium channel activity / ankyrin binding / odontogenesis of dentin-containing tooth / sodium ion transport / regulation of heart rate by cardiac conduction / fibroblast growth factor binding / nitric-oxide synthase binding / intercalated disc / lateral plasma membrane / membrane depolarization / positive regulation of heart rate / neuronal action potential / cardiac muscle contraction / T-tubule / regulation of heart rate / sensory perception of pain / establishment of localization in cell / cellular response to calcium ion / cerebellum development / positive regulation of epithelial cell proliferation / bioluminescence / sodium ion transmembrane transport / generation of precursor metabolites and energy / sarcolemma / caveola / Z disc / scaffold protein binding / transmembrane transporter binding / calmodulin binding / protein domain specific binding / axon / ubiquitin protein ligase binding / protein kinase binding / nucleolus / perinuclear region of cytoplasm / enzyme binding / cell surface / endoplasmic reticulum / nucleoplasm / membrane / plasma membrane Similarity search - Function | |||||||||||||||
| Biological species |   | |||||||||||||||
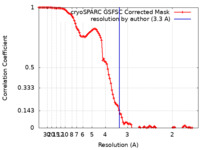
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||||||||
 Authors Authors | Tonggu L / Wisedchaisri G / Gamal El-Din TM / Zheng N / Catterall WA | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Dual receptor-sites reveal the structural basis for hyperactivation of sodium channels by poison-dart toxin batrachotoxin. Authors: Lige Tonggu / Goragot Wisedchaisri / Tamer M Gamal El-Din / Michael J Lenaeus / Matthew M Logan / Tatsuya Toma / Justin Du Bois / Ning Zheng / William A Catterall /   Abstract: The poison dart toxin batrachotoxin is exceptional for its high potency and toxicity, and for its multifaceted modification of the function of voltage-gated sodium channels. By using cryogenic ...The poison dart toxin batrachotoxin is exceptional for its high potency and toxicity, and for its multifaceted modification of the function of voltage-gated sodium channels. By using cryogenic electron microscopy, we identify two homologous, but nonidentical receptor sites that simultaneously bind two molecules of toxin, one at the interface between Domains I and IV, and the other at the interface between Domains III and IV of the cardiac sodium channel. Together, these two bound toxin molecules stabilize α/π helical conformation in the S6 segments that gate the pore, and one of the bound BTX-B molecules interacts with the crucial Lys1421 residue that is essential for sodium conductance and selectivity via an apparent water-bridged hydrogen bond. Overall, our structure provides insight into batrachotoxin's potency, efficacy, and multifaceted functional effects on voltage-gated sodium channels via a dual receptor site mechanism. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41071.map.gz emd_41071.map.gz | 118 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41071-v30.xml emd-41071-v30.xml emd-41071.xml emd-41071.xml | 23.8 KB 23.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41071_fsc.xml emd_41071_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_41071.png emd_41071.png | 143.9 KB | ||
| Filedesc metadata |  emd-41071.cif.gz emd-41071.cif.gz | 8.5 KB | ||
| Others |  emd_41071_half_map_1.map.gz emd_41071_half_map_1.map.gz emd_41071_half_map_2.map.gz emd_41071_half_map_2.map.gz | 116.1 MB 116.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41071 http://ftp.pdbj.org/pub/emdb/structures/EMD-41071 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41071 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41071 | HTTPS FTP |
-Related structure data
| Related structure data |  8t6lMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41071.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41071.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map CryoSPARC | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8255 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map A
| File | emd_41071_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
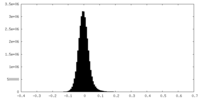
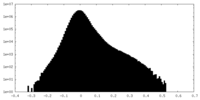
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_41071_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
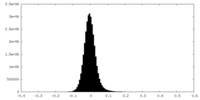
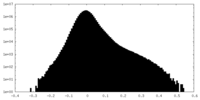
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Sodium channel NaV1.5 and batrachotoxinin-A 20-alpha-benzoate
| Entire | Name: Sodium channel NaV1.5 and batrachotoxinin-A 20-alpha-benzoate |
|---|---|
| Components |
|
-Supramolecule #1: Sodium channel NaV1.5 and batrachotoxinin-A 20-alpha-benzoate
| Supramolecule | Name: Sodium channel NaV1.5 and batrachotoxinin-A 20-alpha-benzoate type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 200 KDa |
-Macromolecule #1: Sodium channel protein type 5 subunit alpha,Green fluorescent protein
| Macromolecule | Name: Sodium channel protein type 5 subunit alpha,Green fluorescent protein type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 213.523875 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MANLLLPRGT SSFRRFTRES LAAIEKRMAE KQTRGGSATS QESREGLQEE EAPRPQLDLQ ASKKLPDLYG NPPRELIGEP LEDLDPFYS TQKTFIVLNK GKTIFRFSAT NALYVLSPFH PVRRAAVKIL VHSLFSMLIM CTILTNCVFM AQHDPPPWTK Y VEYTFTAI ...String: MANLLLPRGT SSFRRFTRES LAAIEKRMAE KQTRGGSATS QESREGLQEE EAPRPQLDLQ ASKKLPDLYG NPPRELIGEP LEDLDPFYS TQKTFIVLNK GKTIFRFSAT NALYVLSPFH PVRRAAVKIL VHSLFSMLIM CTILTNCVFM AQHDPPPWTK Y VEYTFTAI YTFESLVKIL ARGFCLHAFT FLRDPWNWLD FSVIVMAYTT EFVDLDNVSA LRTFRVLRAL KTISVISGLK TI VGALIQS VKKLADVMVL TVFCLSVFAL IGLQLFMGNL RHKCVRNFTE LNGTNGSVEA DGLVWNSLDV YLNDPANYLL KNG TTDVLL CGNSSDAGTC PEGYRCLKAG ENPDHGYTSF DSFAWAFLAL FRLMTQDCWE RLYQQTLRSA GKIYMIFFML VIFL GSFYL VNLILAVVAM AYEEQNQATI AETEEKEKRF QEAMEMLKKE HEALTIRGVD TVSRSSRQRA LSAVSVLTSA LEELE ESHR KCPPCWNRFA QHYLIWECCP LWMSIKQKVK FVVMDPFADL TITMCIVLNT LFMALEHYNM TAEFEEMLQV GNLVFT GIF TAEMTFKIIA LDPYYYFQQG WNIFDSIIVI LSLMELGLSR MGNLSVLRSF RLLRVFKLAK SWPTLNTLIK IIGNSVG AL GNLTLVLAII VFIFAVVGMQ LFGKNYSELR HRISDSGLLP RWHMMDFFHA FLIIFRILCG EWIETMWDCM EVSGQSLC L LVFLLVMVIG NLVVLNLFLA LLLSSFSADN LTAPDEDGEM NNLQLALARI QRGLRFVKRT TWDFCCGILR RRPKKPAAL ATHSQLPSCI TAPRSPPPPE VEKVPPARKE TRFEEDKRPG QGTPGDSEPV CVPIAVAESD TEDQEEDEEN SGKVWWRLRK TCYRIVEHS WFETFIIFMI LLSSGALAFE DIYLEERKTI KVLLEYADKM FTYVFVLEML LKWVAYGFKK YFTNAWCWLD F LIVDVSLV SLVANTLGFA EMGPIKSLRT LRALRPLRAL SRFEGMRVVV NALVGAIPSI MNVLLVCLIF WLIFSIMGVN LF AGKFGRC INQTEGDLPL NYTIVNNKSE CESFNVTGEL YWTKVKVNFD NVGAGYLALL QVATFKGWMD IMYAAVDSRG YEE QPQWED NLYMYIYFVV FIIFGSFFTL NLFIGVIIDN FNQQKKKLGG QDIFMTEEQK KYYNAMKKLG SKKPQKPIPR PLNK YQGFI FDIVTKQAFD VTIMFLICLN MVTMMVETDD QSPEKVNILA KINLLFVAIF TGECIVKMAA LRHYYFTNSW NIFDF VVVI LSIVGTVLSD IIQKYFFSPT LFRVIRLARI GRILRLIRGA KGIRTLLFAL MMSLPALFNI GLLLFLVMFI YSIFGM ANF AYVKWEAGID DMFNFQTFAN SMLCLFQITT SAGWDGLLSP ILNTGPPYCD PNLPNSNGSR GNCGSPAVGI LFFTTYI II SFLIVVNMYI AIILENFSVA TEESTEPLSE DDFDMFYEIW EKFDPEATQF IEYLALSDFA DALSEPLRIA KPNQISLI N MDLPMVSGDR IHCMDILFAF TKRVLGESGE MDALKIQMEE KFMAANPSKI SYEPITTTLR RKHEEVSATV IQRAFRRHL LQRSVKHASF LFRVDLEVLF QGPGSMVSKG EELFTGVVPI LVELDGDVNG HKFSVSGEGE GDATYGKLTL KFICTTGKLP VPWPTLVTT LTYGVQCFSR YPDHMKQHDF FKSAMPEGYV QERTIFFKDD GNYKTRAEVK FEGDTLVNRI ELKGIDFKED G NILGHKLE YNYNSHNVYI MADKQKNGIK VNFKIRHNIE DGSVQLADHY QQNTPIGDGP VLLPDNHYLS TQSALSKDPN EK RDHMVLL EFVTAAGITL GMDELYKGSD YKDDDDK UniProtKB: Sodium channel protein type 5 subunit alpha, Green fluorescent protein |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #6: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine
| Macromolecule | Name: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine / type: ligand / ID: 6 / Number of copies: 10 / Formula: LBN |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-LBN: |
-Macromolecule #7: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 7 / Number of copies: 11 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #8: (3beta,14beta,17beta,25R)-3-[4-methoxy-3-(methoxymethyl)butoxy]sp...
| Macromolecule | Name: (3beta,14beta,17beta,25R)-3-[4-methoxy-3-(methoxymethyl)butoxy]spirost-5-en type: ligand / ID: 8 / Number of copies: 2 / Formula: 9Z9 |
|---|---|
| Molecular weight | Theoretical: 544.805 Da |
| Chemical component information |  ChemComp-9Z9: |
-Macromolecule #9: (1R)-1-[(5aR,7aR,9R,11aS,11bS,12R,13aR)-9,12-dihydroxy-2,11a-dime...
| Macromolecule | Name: (1R)-1-[(5aR,7aR,9R,11aS,11bS,12R,13aR)-9,12-dihydroxy-2,11a-dimethyl-1,2,3,4,7a,8,9,10,11,11a,12,13-dodecahydro-7H-9,11b-epoxy-13a,5a-prop[1]enophenanthro[2,1-f][1,4]oxazepin-14-yl]ethyl benzoate type: ligand / ID: 9 / Number of copies: 2 / Formula: YIJ |
|---|---|
| Molecular weight | Theoretical: 521.644 Da |
-Macromolecule #10: water
| Macromolecule | Name: water / type: ligand / ID: 10 / Number of copies: 1 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 6 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 2 / Number real images: 7542 / Average exposure time: 2.04 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 49.4 / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-8t6l: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)