+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of NKCC1 Fu_CTD | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cotransporter / dimer / ion / membrane protein / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsodium:potassium:chloride symporter activity / sodium ion homeostasis / ammonium channel activity / chloride ion homeostasis / potassium ion homeostasis / cell volume homeostasis / potassium ion import across plasma membrane / apical plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
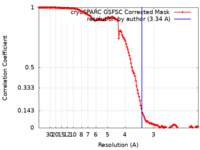
| Method | single particle reconstruction / cryo EM / Resolution: 3.34 Å | |||||||||
 Authors Authors | Moseng MA | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Inhibition mechanism of NKCC1 involves the carboxyl terminus and long-range conformational coupling. Authors: Mitchell A Moseng / Chih-Chia Su / Kerri Rios / Meng Cui / Meinan Lyu / Przemyslaw Glaza / Philip A Klenotic / Eric Delpire / Edward W Yu /  Abstract: The Na-K-2Cl cotransporter-1 (NKCC1) is an electroneutral Na-dependent transporter responsible for simultaneously translocating Na, K, and Cl ions into cells. In human tissue, NKCC1 plays a critical ...The Na-K-2Cl cotransporter-1 (NKCC1) is an electroneutral Na-dependent transporter responsible for simultaneously translocating Na, K, and Cl ions into cells. In human tissue, NKCC1 plays a critical role in regulating cytoplasmic volume, fluid intake, chloride homeostasis, and cell polarity. Here, we report four structures of human NKCC1 (hNKCC1), both in the absence and presence of loop diuretic (bumetanide or furosemide), using single-particle cryo-electron microscopy. These structures allow us to directly observe various novel conformations of the hNKCC1 dimer. They also reveal two drug-binding sites located at the transmembrane and cytosolic carboxyl-terminal domains, respectively. Together, our findings enable us to delineate an inhibition mechanism that involves a coupled movement between the cytosolic and transmembrane domains of hNKCC1. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40759.map.gz emd_40759.map.gz | 230.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40759-v30.xml emd-40759-v30.xml emd-40759.xml emd-40759.xml | 13.6 KB 13.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40759_fsc.xml emd_40759_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_40759.png emd_40759.png | 108.3 KB | ||
| Filedesc metadata |  emd-40759.cif.gz emd-40759.cif.gz | 5.4 KB | ||
| Others |  emd_40759_half_map_1.map.gz emd_40759_half_map_1.map.gz emd_40759_half_map_2.map.gz emd_40759_half_map_2.map.gz | 226.7 MB 226.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40759 http://ftp.pdbj.org/pub/emdb/structures/EMD-40759 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40759 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40759 | HTTPS FTP |
-Related structure data
| Related structure data |  8steMC  7mxoC  7n3nC  7sflC  7smpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40759.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40759.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_40759_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
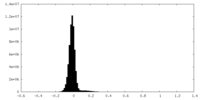
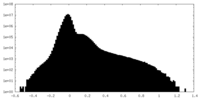
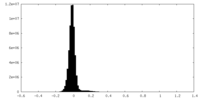
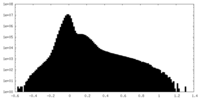
| Density Histograms |
-Half map: #1
| File | emd_40759_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : NKCC1_Fu_CTD
| Entire | Name: NKCC1_Fu_CTD |
|---|---|
| Components |
|
-Supramolecule #1: NKCC1_Fu_CTD
| Supramolecule | Name: NKCC1_Fu_CTD / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Solute carrier family 12 member 2
| Macromolecule | Name: Solute carrier family 12 member 2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.190438 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SSTQALTYLN ALQHSIRLSG VEDHVKNFRP QCLVMTGAPN SRPALLHLVH DFTKNVGLMI CGHVHMGPRR QAMKEMSIDQ AKYQRWLIK NKMKAFYAPV HADDLREGAQ YLMQAAGLGR MKPNTLVLGF KKDWLQADMR DVDMYINLFH DAFDIQYGVV V IRLKEGLD ...String: SSTQALTYLN ALQHSIRLSG VEDHVKNFRP QCLVMTGAPN SRPALLHLVH DFTKNVGLMI CGHVHMGPRR QAMKEMSIDQ AKYQRWLIK NKMKAFYAPV HADDLREGAQ YLMQAAGLGR MKPNTLVLGF KKDWLQADMR DVDMYINLFH DAFDIQYGVV V IRLKEGLD ISHLQGQESK GPIVPLNVAD QKLLEASTQF QKKQGKNTID VWWLFDDGGL TLLIPYLLTT KKKWKDCKIR VF IGGKINR IDHDRRAMAT LLSKFRIDFS DIMVLGDINT KPKKENIIAF EEIIEPYRLH EDDKEQDIAD KMKEDEPWRI TDN ELELYK TKTYRQIRLN ELLKEHSSTA NIIVMSLPVA RKGAVSSALY MAWLEALSKD LPPILLVRGN HQSVLTF UniProtKB: Solute carrier family 12 member 2 |
-Macromolecule #2: 5-(AMINOSULFONYL)-4-CHLORO-2-[(2-FURYLMETHYL)AMINO]BENZOIC ACID
| Macromolecule | Name: 5-(AMINOSULFONYL)-4-CHLORO-2-[(2-FURYLMETHYL)AMINO]BENZOIC ACID type: ligand / ID: 2 / Number of copies: 2 / Formula: FUN |
|---|---|
| Molecular weight | Theoretical: 330.744 Da |
| Chemical component information | 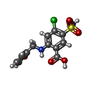 ChemComp-FUN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)