[English] 日本語
 Yorodumi
Yorodumi- EMDB-40753: The 2alpha3beta stoichiometry of full-length human alpha4beta2 ni... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | The 2alpha3beta stoichiometry of full-length human alpha4beta2 nicotinic acetylcholine receptor in complex with acetylcholine | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | cys-loop ligand-gated pentameric ion channels / cation-selective channel / acetylcholine / TRANSPORT PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationvestibulocochlear nerve development / lateral geniculate nucleus development / regulation of circadian sleep/wake cycle, REM sleep / regulation of synaptic transmission, dopaminergic / optic nerve morphogenesis / Highly sodium permeable postsynaptic acetylcholine nicotinic receptors / Highly calcium permeable nicotinic acetylcholine receptors / central nervous system projection neuron axonogenesis / synaptic transmission involved in micturition / response to acetylcholine ...vestibulocochlear nerve development / lateral geniculate nucleus development / regulation of circadian sleep/wake cycle, REM sleep / regulation of synaptic transmission, dopaminergic / optic nerve morphogenesis / Highly sodium permeable postsynaptic acetylcholine nicotinic receptors / Highly calcium permeable nicotinic acetylcholine receptors / central nervous system projection neuron axonogenesis / synaptic transmission involved in micturition / response to acetylcholine / negative regulation of action potential / Highly calcium permeable postsynaptic nicotinic acetylcholine receptors / acetylcholine receptor activity / acetylcholine-gated channel complex / behavioral response to nicotine / regulation of dopamine metabolic process / positive regulation of dopamine secretion / neuromuscular synaptic transmission / acetylcholine-gated monoatomic cation-selective channel activity / cation channel complex / acetylcholine binding / synaptic transmission, cholinergic / nervous system process / neurotransmitter receptor complex / ATP-dependent chromatin remodeler activity / acetylcholine receptor signaling pathway / regulation of dendrite morphogenesis / inhibitory postsynaptic potential / regulation of synapse assembly / regulation of dopamine secretion / B cell activation / smooth muscle contraction / action potential / associative learning / membrane depolarization / plasma membrane raft / social behavior / ligand-gated monoatomic ion channel activity / positive regulation of B cell proliferation / monoatomic ion transport / visual perception / sensory perception of pain / presynaptic modulation of chemical synaptic transmission / learning / response to nicotine / response to cocaine / locomotory behavior / regulation of membrane potential / sensory perception of sound / visual learning / cognition / memory / calcium ion transport / response to oxidative stress / presynaptic membrane / monoatomic ion transmembrane transport / chemical synaptic transmission / response to ethanol / response to hypoxia / postsynaptic membrane / neuron projection / external side of plasma membrane / DNA repair / neuronal cell body / synapse / dendrite / signal transduction / membrane / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
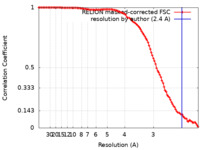
| Method | single particle reconstruction / cryo EM / Resolution: 2.4 Å | ||||||||||||
 Authors Authors | Kang G / Hibbs RE | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Br J Pharmacol / Year: 2024 Journal: Br J Pharmacol / Year: 2024Title: Structural bases for stoichiometry-selective calcium potentiation of a neuronal nicotinic receptor. Authors: Simone Mazzaferro / Guipeun Kang / Kathiresan Natarajan / Ryan E Hibbs / Steven M Sine /   Abstract: BACKGROUND AND PURPOSE: α4β2 nicotinic acetylcholine (nACh) receptors assemble in two stoichiometric forms, one of which is potentiated by calcium. The sites of calcium binding that underpin potentiation are not known. EXPERIMENTAL APPROACH: To identify calcium binding sites, we applied cryo-electron microscopy (cryo-EM) and molecular dynamics (MD) simulations to each stoichiometric form of the α4β2 nACh receptor ...EXPERIMENTAL APPROACH: To identify calcium binding sites, we applied cryo-electron microscopy (cryo-EM) and molecular dynamics (MD) simulations to each stoichiometric form of the α4β2 nACh receptor in the presence of calcium ions. To test whether the identified calcium sites are linked to potentiation, we generated mutants of anionic residues at the sites, expressed wild type and mutant receptors in clonal mammalian fibroblasts, and recorded ACh-elicited single-channel currents with or without calcium. KEY RESULTS: Both cryo-EM and MD simulations show calcium bound to a site between the extracellular and transmembrane domains of each α4 subunit (ECD-TMD site). Substituting alanine for anionic ...KEY RESULTS: Both cryo-EM and MD simulations show calcium bound to a site between the extracellular and transmembrane domains of each α4 subunit (ECD-TMD site). Substituting alanine for anionic residues at the ECD-TMD site abolishes stoichiometry-selective calcium potentiation, as monitored by single-channel patch clamp electrophysiology. Additionally, MD simulation reveals calcium association at subunit interfaces within the extracellular domain. Substituting alanine for anionic residues at the ECD sites reduces or abolishes stoichiometry-selective calcium potentiation. CONCLUSIONS AND IMPLICATIONS: Stoichiometry-selective calcium potentiation of the α4β2 nACh receptor is achieved by calcium association with topographically distinct sites framed by anionic ...CONCLUSIONS AND IMPLICATIONS: Stoichiometry-selective calcium potentiation of the α4β2 nACh receptor is achieved by calcium association with topographically distinct sites framed by anionic residues within the α4 subunit and between the α4 and β2 subunits. Stoichiometry-selective calcium potentiation could result from the greater number of calcium sites in the stoichiometric form with three rather than two α4 subunits. The results are relevant to modulation of signalling via α4β2 nACh receptors in physiological and pathophysiological conditions. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40753.map.gz emd_40753.map.gz | 11.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40753-v30.xml emd-40753-v30.xml emd-40753.xml emd-40753.xml | 25.3 KB 25.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40753_fsc.xml emd_40753_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_40753.png emd_40753.png | 442.3 KB | ||
| Filedesc metadata |  emd-40753.cif.gz emd-40753.cif.gz | 8 KB | ||
| Others |  emd_40753_half_map_1.map.gz emd_40753_half_map_1.map.gz emd_40753_half_map_2.map.gz emd_40753_half_map_2.map.gz | 80.5 MB 80.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40753 http://ftp.pdbj.org/pub/emdb/structures/EMD-40753 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40753 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40753 | HTTPS FTP |
-Related structure data
| Related structure data |  8st0MC  8sszC  8st1C  8st2C  8st3C  8st4C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40753.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40753.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.079 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_40753_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_40753_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : A complex of three Fab fragments with the 2alpha3beta stoichiomet...
+Supramolecule #1: A complex of three Fab fragments with the 2alpha3beta stoichiomet...
+Macromolecule #1: Neuronal acetylcholine receptor subunit alpha-4
+Macromolecule #2: Neuronal acetylcholine receptor subunit beta-2
+Macromolecule #3: IgG1 Kappa Light Chain
+Macromolecule #4: IgG1 Heavy Chain
+Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
+Macromolecule #7: ACETYLCHOLINE
+Macromolecule #8: N-OCTANE
+Macromolecule #9: SODIUM ION
+Macromolecule #10: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)