+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Phosphoinositide phosphate 3 kinase gamma | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Phosphoinositide 3-Kinase / Phosphoinositide 3-Kinase /  Chemotaxis / Chemotaxis /  Cancer / Cancer /  SIGNALING PROTEIN SIGNALING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information1-phosphatidylinositol-3-kinase regulator activity / phosphatidylinositol 3-kinase complex, class IB /  phosphatidylinositol 3-kinase complex / 1-phosphatidylinositol-4-phosphate 3-kinase activity / phosphatidylinositol 3-kinase complex / 1-phosphatidylinositol-4-phosphate 3-kinase activity /  phosphatidylinositol-4,5-bisphosphate 3-kinase / 1-phosphatidylinositol-3-kinase activity / phosphatidylinositol-mediated signaling / phosphatidylinositol-4,5-bisphosphate 3-kinase / 1-phosphatidylinositol-3-kinase activity / phosphatidylinositol-mediated signaling /  cell migration / G protein-coupled receptor signaling pathway / cell migration / G protein-coupled receptor signaling pathway /  phosphorylation ...1-phosphatidylinositol-3-kinase regulator activity / phosphatidylinositol 3-kinase complex, class IB / phosphorylation ...1-phosphatidylinositol-3-kinase regulator activity / phosphatidylinositol 3-kinase complex, class IB /  phosphatidylinositol 3-kinase complex / 1-phosphatidylinositol-4-phosphate 3-kinase activity / phosphatidylinositol 3-kinase complex / 1-phosphatidylinositol-4-phosphate 3-kinase activity /  phosphatidylinositol-4,5-bisphosphate 3-kinase / 1-phosphatidylinositol-3-kinase activity / phosphatidylinositol-mediated signaling / phosphatidylinositol-4,5-bisphosphate 3-kinase / 1-phosphatidylinositol-3-kinase activity / phosphatidylinositol-mediated signaling /  cell migration / G protein-coupled receptor signaling pathway / cell migration / G protein-coupled receptor signaling pathway /  phosphorylation / phosphorylation /  nucleus / nucleus /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Sus scrofa (pig) Sus scrofa (pig) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.03 Å cryo EM / Resolution: 3.03 Å | |||||||||
 Authors Authors | Chen C-L / Tesmer JJG / Bandekar SJ / Cash J | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Molecular basis for Gβγ-mediated activation of phosphoinositide 3-kinase γ. Authors: Chun-Liang Chen / Ramizah Syahirah / Sandeep K Ravala / Yu-Chen Yen / Thomas Klose / Qing Deng / John J G Tesmer /  Abstract: The conversion of phosphatidylinositol 4,5-bisphosphate to phosphatidylinositol 3,4,5-triphosphate by phosphoinositide 3-kinase γ (PI3Kγ) is critical for neutrophil chemotaxis and cancer metastasis. ...The conversion of phosphatidylinositol 4,5-bisphosphate to phosphatidylinositol 3,4,5-triphosphate by phosphoinositide 3-kinase γ (PI3Kγ) is critical for neutrophil chemotaxis and cancer metastasis. PI3Kγ is activated by Gβγ heterodimers released from G protein-coupled receptors responding to extracellular signals. Here we determined cryo-electron microscopy structures of Sus scrofa PI3Kγ-human Gβγ complexes in the presence of substrates/analogs, revealing two Gβγ binding sites: one on the p110γ helical domain and another on the p101 C-terminal domain. Comparison with PI3Kγ alone reveals conformational changes in the kinase domain upon Gβγ binding that are similar to Ras·GTP-induced changes. Assays of variants perturbing the Gβγ binding sites and interdomain contacts altered by Gβγ binding suggest that Gβγ recruits the enzyme to membranes and allosterically regulates activity via both sites. Studies of zebrafish neutrophil migration align with these findings, paving the way for in-depth investigation of Gβγ-mediated activation mechanisms in this enzyme family and drug development for PI3Kγ. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40650.map.gz emd_40650.map.gz | 114.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40650-v30.xml emd-40650-v30.xml emd-40650.xml emd-40650.xml | 23 KB 23 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40650_fsc.xml emd_40650_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_40650.png emd_40650.png | 91 KB | ||
| Filedesc metadata |  emd-40650.cif.gz emd-40650.cif.gz | 7.7 KB | ||
| Others |  emd_40650_half_map_1.map.gz emd_40650_half_map_1.map.gz emd_40650_half_map_2.map.gz emd_40650_half_map_2.map.gz | 59.5 MB 59.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40650 http://ftp.pdbj.org/pub/emdb/structures/EMD-40650 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40650 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40650 | HTTPS FTP |
-Related structure data
| Related structure data |  8so9MC  8soaC  8sobC  8socC  8sodC  8soeC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40650.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40650.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_40650_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
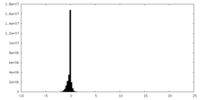
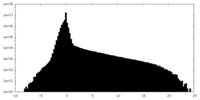
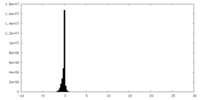
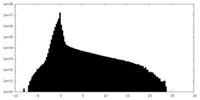
| Density Histograms |
-Half map: #1
| File | emd_40650_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PI3K-gamma containing the p110gamma catalytic subunit and the p10...
| Entire | Name: PI3K-gamma containing the p110gamma catalytic subunit and the p101 regulatory subunit |
|---|---|
| Components |
|
-Supramolecule #1: PI3K-gamma containing the p110gamma catalytic subunit and the p10...
| Supramolecule | Name: PI3K-gamma containing the p110gamma catalytic subunit and the p101 regulatory subunit type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Sus scrofa (pig) Sus scrofa (pig) |
| Molecular weight | Theoretical: 210 kDa/nm |
-Macromolecule #1: phosphatidylinositol-4,5-bisphosphate 3-kinase
| Macromolecule | Name: phosphatidylinositol-4,5-bisphosphate 3-kinase / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Sus scrofa (pig) Sus scrofa (pig) |
| Molecular weight | Theoretical: 127.573531 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MHHHHHHELE NYEQPVVLRE DNRRRRRRMK PRSTAASLSS MELIPIEFVL PTSQRNTKTP ETALLHVAGH GNVEQMKAQV WLRALETSV SADFYHRLGP DHFLLLYQKK GQWYEIYDKY QVVQTLDCLR YWKVLHRSPG QIHVVQRHAP SEETLAFQRQ L NALIGYDV ...String: MHHHHHHELE NYEQPVVLRE DNRRRRRRMK PRSTAASLSS MELIPIEFVL PTSQRNTKTP ETALLHVAGH GNVEQMKAQV WLRALETSV SADFYHRLGP DHFLLLYQKK GQWYEIYDKY QVVQTLDCLR YWKVLHRSPG QIHVVQRHAP SEETLAFQRQ L NALIGYDV TDVSNVHDDE LEFTRRRLVT PRMAEVAGRD PKLYAMHPWV TSKPLPEYLL KKITNNCVFI VIHRSTTSQT IK VSADDTP GTILQSFFTK MAKKKSLMDI PESQNERDFV LRVCGRDEYL VGETPIKNFQ WVRQCLKNGE EIHLVLDTPP DPA LDEVRK EEWPLVDDCT GVTGYHEQLT IHGKDHESVF TVSLWDCDRK FRVKIRGIDI PVLPRTADLT VFVEANIQYG QQVL CQRRT SPKPFTEEVL WNVWLEFSIK IKDLPKGALL NLQIYCGKAP ALSGKTSAEM PSPESKGKAQ LLYYVNLLLI DHRFL LRHG EYVLHMWQLS GKGEDQGSFN ADKLTSATNP DKENSMSISI LLDNYCHPIA LPKHRPTPDP EGDRVRAEMP NQLRKQ LEA IIATDPLNPL TAEDKELLWH FRYESLKDPK AYPKLFSSVK WGQQEIVAKT YQLLAKREVW DQSALDVGLT MQLLDCN FS DENVRAIAVQ KLESLEDDDV LHYLLQLVQA VKFEPYHDSA LARFLLKRGL RNKRIGHFLF WFLRSEIAQS RHYQQRFA V ILEAYLRGCG TAMLHDFTQQ VQVIDMLQKV TIDIKSLSAE KYDVSSQVIS QLKQKLENLQ NLNLPQSFRV PYDPGLKAG ALVIEKCKVM ASKKKPLWLE FKCADPTALS NETIGIIFKH GDDLRQDMLI LQILRIMESI WETESLDLCL LPYGCISTGD KIGMIEIVK DATTIAKIQQ STVGNTGAFK DEVLSHWLKE KCPIEEKFQA AVERFVYSCA GYCVATFVLG IGDRHNDNIM I SETGNLFH IDFGHILGNY KSFLGINKER VPFVLTPDFL FVMGTSGKKT SLHFQKFQDV CVKAYLALRH HTNLLIILFS MM LMTGMPQ LTSKEDIEYI RDALTVGKSE EDAKKYFLDQ IEVCRDKGWT VQFNWFLHLV LGIKQGEKHS A UniProtKB:  phosphatidylinositol-4,5-bisphosphate 3-kinase phosphatidylinositol-4,5-bisphosphate 3-kinase |
-Macromolecule #2: Phosphoinositide 3-kinase regulatory subunit 5
| Macromolecule | Name: Phosphoinositide 3-kinase regulatory subunit 5 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Sus scrofa (pig) Sus scrofa (pig) |
| Molecular weight | Theoretical: 98.497773 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MQPGATTCTE DRIQHALERC LHGLSLSRRS TSWSAGLCLN CWSLQELVSR DPGHFLILLE QILQKTREVQ EKGTYDLLAP LALLFYSTV LCTPHFPPDS DLLLKAARTY HRFLTWPVPY CSICQELLTF IDAELKAPGI SYQRLVRAEQ GLSTRSHRSS T VTVLLLNP ...String: MQPGATTCTE DRIQHALERC LHGLSLSRRS TSWSAGLCLN CWSLQELVSR DPGHFLILLE QILQKTREVQ EKGTYDLLAP LALLFYSTV LCTPHFPPDS DLLLKAARTY HRFLTWPVPY CSICQELLTF IDAELKAPGI SYQRLVRAEQ GLSTRSHRSS T VTVLLLNP VEVQAEFLDV ADKLSTPGPS PHSAYITLLL HAFQATFGAH CDLSGLHRRL QSKTLAELEA IFTETAEAQE LA SGIGDAA EARQWLRTKL QAVGEKAGFP GVLDTAKPGK LRTIPIPVAR CYTYSWNQDS FDILQEILLK EQELLQPEIL DDE EDEDEE DEEEDLDADG HCAERDSVLS TGSAASHAST LSLASSQASG PTLSRQLLTS FVSGLSDGVD SGYMEDIEES AYER PRRPG GHERRGHRRP GQKFNRIYKL FKSTSQMVLR RDSRSLEGSP DSGPPLRRAG SLCSPLDSPT LPPSRAQRSR SLPQP KLSP QLPGWLLAPA SRHQRRRPFL SGDEDPKAST LRVVVFGSDR ISGKVARAYS NLRRLENNRP LLTRFFKLQF FYVPVK RSR GTGTPTSPAP RSQTPPLPTD APRHPGPAEL GAAPWEESTN DISHYLGMLD PWYERNVLGL MHLPPEVLCQ SLKAEPR PL EGSPAQLPIL ADMLLYYCRF AARPVLLQVY QTELTFITGE KTTEIFIHSL ELGHSAATRA IKASGPGSKR LGIDGDRE A VPLTLQIIYS KGAISGRSRW SNMEKLCTSV NLSKACRQQE ELDSSTEALT LNLTEVVKRQ TPKSKKGFNQ ISTSQIKVD KVQIIGSNSC PFAVCLDQDE RKILQSVIRC EVSPCYKPEK SSLCPPPQRP SYPPAPATPD LCSLLCLPIM TFSGALPGGG GSDYKDDDD K UniProtKB: Phosphoinositide 3-kinase regulatory subunit 5 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Details: Glow discharge with 20 mA for 1 minute | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: Blot force 2. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Specialist optics | Energy filter - Name: GIF Quantum ER / Energy filter - Slit width: 20 eV |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Alignment procedure | Coma free - Residual tilt: 0.01 mrad |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 11520 pixel / Digitization - Dimensions - Height: 8184 pixel / Average exposure time: 3.12 sec. / Average electron dose: 55.0 e/Å2 Details: Images were collected in movie-mode at 40 frames per second |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Target criteria: correlation coefficient | ||||||
| Output model |  PDB-8so9: |
 Movie
Movie Controller
Controller












 Z
Z Y
Y X
X