+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of PAPP-A2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Protease / zinc binding / growth factor signaling / peptide binding / HYDROLASE / PEPTIDE BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationHydrolases; Acting on peptide bonds (peptidases); Metalloendopeptidases / bone morphogenesis / response to salt stress / regulation of cell growth / metalloendopeptidase activity / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / metallopeptidase activity / cell surface receptor signaling pathway / apical plasma membrane / proteolysis ...Hydrolases; Acting on peptide bonds (peptidases); Metalloendopeptidases / bone morphogenesis / response to salt stress / regulation of cell growth / metalloendopeptidase activity / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / metallopeptidase activity / cell surface receptor signaling pathway / apical plasma membrane / proteolysis / : / extracellular exosome / extracellular region / zinc ion binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
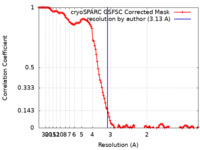
| Method | single particle reconstruction / cryo EM / Resolution: 3.13 Å | |||||||||
 Authors Authors | Judge RA / Stoll VS / Eaton D / Hao Q / Bratkowski MA | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Commun Chem / Year: 2023 Journal: Commun Chem / Year: 2023Title: Cryo-EM structure of human PAPP-A2 and mechanism of substrate recognition. Authors: Janani Sridar / Amirhossein Mafi / Russell A Judge / Jun Xu / Kailyn A Kong / John C K Wang / Vincent S Stoll / Georgios Koukos / Reyna J Simon / Dan Eaton / Matthew Bratkowski / Qi Hao /  Abstract: Pregnancy-Associated Plasma Protein A isoforms, PAPP-A and PAPP-A2, are metalloproteases that cleave insulin-like growth factor binding proteins (IGFBPs) to modulate insulin-like growth factor ...Pregnancy-Associated Plasma Protein A isoforms, PAPP-A and PAPP-A2, are metalloproteases that cleave insulin-like growth factor binding proteins (IGFBPs) to modulate insulin-like growth factor signaling. The structures of homodimeric PAPP-A in complex with IGFBP5 anchor peptide, and inhibitor proteins STC2 and proMBP have been recently reported. Here, we present the single-particle cryo-EM structure of the monomeric, N-terminal LG, MP, and the M1 domains (with the exception of LNR1/2) of human PAPP-A2 to 3.13 Å resolution. Our structure together with functional studies provides insight into a previously reported patient mutation that inactivates PAPP-A2 in a distal region of the protein. Using a combinational approach, we suggest that PAPP-A2 recognizes IGFBP5 in a similar manner as PAPP-A and show that PAPP-A2 cleaves IGFBP5 less efficiently due to differences in the M2 domain. Overall, our studies characterize the cleavage mechanism of IGFBP5 by PAPP-A2 and shed light onto key differences with its paralog PAPP-A. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40571.map.gz emd_40571.map.gz | 167.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40571-v30.xml emd-40571-v30.xml emd-40571.xml emd-40571.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40571_fsc.xml emd_40571_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_40571.png emd_40571.png | 46 KB | ||
| Masks |  emd_40571_msk_1.map emd_40571_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-40571.cif.gz emd-40571.cif.gz | 7.5 KB | ||
| Others |  emd_40571_half_map_1.map.gz emd_40571_half_map_1.map.gz emd_40571_half_map_2.map.gz emd_40571_half_map_2.map.gz | 165 MB 165 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40571 http://ftp.pdbj.org/pub/emdb/structures/EMD-40571 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40571 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40571 | HTTPS FTP |
-Related structure data
| Related structure data |  8sl1MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40571.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40571.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.669 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40571_msk_1.map emd_40571_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
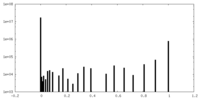
| Density Histograms |
-Half map: #2
| File | emd_40571_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
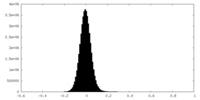
| Density Histograms |
-Half map: #1
| File | emd_40571_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
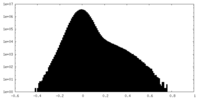
| Density Histograms |
- Sample components
Sample components
-Entire : PAPP-A2
| Entire | Name: PAPP-A2 |
|---|---|
| Components |
|
-Supramolecule #1: PAPP-A2
| Supramolecule | Name: PAPP-A2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Pappalysin-2
| Macromolecule | Name: Pappalysin-2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on peptide bonds (peptidases); Metalloendopeptidases |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 174.259766 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: PPEESNQNGG EGSYREAETF NSQVGLPILY FSGRRERLLL RPEVLAEIPR EAFTVEAWVK PEGGQNNPAI IAGVFDNCSH TVSDKGWAL GIRSGKDKGK RDARFFFSLC TDRVKKATIL ISHSRYQPGT WTHVAATYDG RHMALYVDGT QVASSLDQSG P LNSPFMAS ...String: PPEESNQNGG EGSYREAETF NSQVGLPILY FSGRRERLLL RPEVLAEIPR EAFTVEAWVK PEGGQNNPAI IAGVFDNCSH TVSDKGWAL GIRSGKDKGK RDARFFFSLC TDRVKKATIL ISHSRYQPGT WTHVAATYDG RHMALYVDGT QVASSLDQSG P LNSPFMAS CRSLLLGGDS SEDGHYFRGH LGTLVFWSTA LPQSHFQHSS QHSSGEEEAT DLVLTASFEP VNTEWVPFRD EK YPRLEVL QGFEPEPEIL SPLQPPLCGQ TVCDNVELIS QYNGYWPLRG EKVIRYQVVN ICDDEGLNPI VSEEQIRLQH EAL NEAFSR YNISWQLSVH QVHNSTLRHR VVLVNCEPSK IGNDHCDPEC EHPLTGYDGG DCRLQGRCYS WNRRDGLCHV ECNN MLNDF DDGDCCDPQV ADVRKTCFDP DSPKRAYMSV KELKEALQLN STHFLNIYFA SSVREDLAGA ATWPWDKDAV THLGG IVLS PAYYGMPGHT DTMIHQVGHV LGLYHVFKGV SERESCNDPC KETVPSMETG DLCADTAPTP KSELCREPEP TSDTCG FTR FPGAPFTNYM SYTDDNCTDN FTPNQVARMH CYLDLVYQQW TESRKPTPIP IPPMVIGQTN KSLTIHWLPP ISGVVYD RA SGSLCGACTE DGTFRQYVHT ASSRRVCDSS GYWTPEEAVG PPDVDQPCEP SLQAWSPEVH LYHMNMTVPC PTEGCSLE L LFQHPVQADT LTLWVTSFFM ESSQVLFDTE ILLENKESVH LGPLDTFCDI PLTIKLHVDG KVSGVKVYTF DERIEIDAA LLTSQPHSPL CSGCRPVRYQ VLRDPPFASG LPVVVTHSHR KFTDVEVTPG QMYQYQVLAE AGGELGEASP PLNHIHGAPY CGDGKVSER LGEECDDGDL VSGDGCSKVC ELEEGFNCVG EPSLCYMYEG DGICEPFERK TSIVDCGIYT PKGYLDQWAT R AYSSHEDK KKCPVSLVTG EPHSLICTSY HPDLPNHRPL TGWFPCVASE NETQDDRSEQ PEGSLKKEDE VWLKVCFNRP GE ARAIFIF LTTDGLVPGE HQQPTVTLYL TDVRGSNHSL GTYGLSCQHN PLIINVTHHQ NVLFHHTTSV LLNFSSPRVG ISA VALRTS SRIGLSAPSN CISEDEGQNH QGQSCIHRPC GKQDSCPSLL LDHADVVNCT SIGPGLMKCA ITCQRGFALQ ASSG QYIRP MQKEILLTCS SGHWDQNVSC LPVDCGVPDP SLVNYANFSC SEGTKFLKRC SISCVPPAKL QGLSPWLTCL EDGLW SLPE VYCKLECDAP PIILNANLLL PHCLQDNHDV GTICKYECKP GYYVAESAEG KVRNKLLKIQ CLEGGIWEQG SCIPVV CEP PPPVFEGMYE CTNGFSLDSQ CVLNCNQERE KLPILCTKEG LWTQEFKLCE NLQGECPPPP SELNSVEYKC EQGYGIG AV CSPLCVIPPS DPVMLPENIT ADTLEHWMEP VKVQSIVCTG RRQWHPDPVL VHCIQSCEPF QADGWCDTIN NRAYCHYD G GDCCSSTLSS KKVIPFAADC DLDECTCRDP KAEENQTRTG GDYKDDDDK UniProtKB: Pappalysin-2 |
-Macromolecule #3: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 1 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.4000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)