+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human NCX1 in apo inactivated state | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Na/Ca exchanger / sodium calcium exchanger / TRANSPORT PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationrelaxation of smooth muscle / vascular associated smooth muscle contraction / negative regulation of intracellular signal transduction / calcium:sodium antiporter activity / regulation of cell communication by electrical coupling / calcium ion export / membrane depolarization during cardiac muscle cell action potential / regulation of the force of heart contraction / cell communication by electrical coupling involved in cardiac conduction / sodium ion export across plasma membrane ...relaxation of smooth muscle / vascular associated smooth muscle contraction / negative regulation of intracellular signal transduction / calcium:sodium antiporter activity / regulation of cell communication by electrical coupling / calcium ion export / membrane depolarization during cardiac muscle cell action potential / regulation of the force of heart contraction / cell communication by electrical coupling involved in cardiac conduction / sodium ion export across plasma membrane / intracellular sodium ion homeostasis / sodium ion import across plasma membrane / calcium ion import / calcium ion transport into cytosol / relaxation of cardiac muscle / cardiac muscle cell development / regulation of cardiac muscle contraction by calcium ion signaling / Sodium/Calcium exchangers / ankyrin binding / Reduction of cytosolic Ca++ levels / cellular response to caffeine / negative regulation of cytosolic calcium ion concentration / calcium ion transmembrane import into cytosol / positive regulation of the force of heart contraction / calcium ion import across plasma membrane / intercalated disc / regulation of cardiac conduction / positive regulation of bone mineralization / regulation of cardiac muscle contraction by regulation of the release of sequestered calcium ion / calcium ion homeostasis / cytoskeletal protein binding / cardiac muscle contraction / Ion homeostasis / monoatomic ion transport / response to muscle stretch / axon terminus / T-tubule / sodium ion transmembrane transport / muscle contraction / regulation of heart rate / cell periphery / cellular response to reactive oxygen species / sarcolemma / calcium ion transmembrane transport / Z disc / intracellular calcium ion homeostasis / regulation of gene expression / transmembrane transporter binding / calmodulin binding / postsynapse / postsynaptic density / axon / neuronal cell body / dendrite / calcium ion binding / synapse / nucleoplasm / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
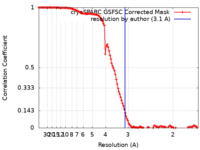
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | ||||||||||||
 Authors Authors | Xue J / Jiang Y | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural mechanisms of the human cardiac sodium-calcium exchanger NCX1. Authors: Jing Xue / Weizhong Zeng / Yan Han / Scott John / Michela Ottolia / Youxing Jiang /  Abstract: Na/Ca exchangers (NCX) transport Ca in or out of cells in exchange for Na. They are ubiquitously expressed and play an essential role in maintaining cytosolic Ca homeostasis. Although extensively ...Na/Ca exchangers (NCX) transport Ca in or out of cells in exchange for Na. They are ubiquitously expressed and play an essential role in maintaining cytosolic Ca homeostasis. Although extensively studied, little is known about the global structural arrangement of eukaryotic NCXs and the structural mechanisms underlying their regulation by various cellular cues including cytosolic Na and Ca. Here we present the cryo-EM structures of human cardiac NCX1 in both inactivated and activated states, elucidating key structural elements important for NCX ion exchange function and its modulation by cytosolic Ca and Na. We demonstrate that the interactions between the ion-transporting transmembrane (TM) domain and the cytosolic regulatory domain define the activity of NCX. In the inward-facing state with low cytosolic [Ca], a TM-associated four-stranded β-hub mediates a tight packing between the TM and cytosolic domains, resulting in the formation of a stable inactivation assembly that blocks the TM movement required for ion exchange function. Ca binding to the cytosolic second Ca-binding domain (CBD2) disrupts this inactivation assembly which releases its constraint on the TM domain, yielding an active exchanger. Thus, the current NCX1 structures provide an essential framework for the mechanistic understanding of the ion transport and cellular regulation of NCX family proteins. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40457.map.gz emd_40457.map.gz | 167.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40457-v30.xml emd-40457-v30.xml emd-40457.xml emd-40457.xml | 26.2 KB 26.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40457_fsc.xml emd_40457_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_40457.png emd_40457.png | 74.2 KB | ||
| Filedesc metadata |  emd-40457.cif.gz emd-40457.cif.gz | 7.6 KB | ||
| Others |  emd_40457_additional_1.map.gz emd_40457_additional_1.map.gz emd_40457_additional_2.map.gz emd_40457_additional_2.map.gz emd_40457_half_map_1.map.gz emd_40457_half_map_1.map.gz emd_40457_half_map_2.map.gz emd_40457_half_map_2.map.gz | 167.9 MB 167.8 MB 165.3 MB 165.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40457 http://ftp.pdbj.org/pub/emdb/structures/EMD-40457 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40457 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40457 | HTTPS FTP |
-Validation report
| Summary document |  emd_40457_validation.pdf.gz emd_40457_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40457_full_validation.pdf.gz emd_40457_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_40457_validation.xml.gz emd_40457_validation.xml.gz | 20.3 KB | Display | |
| Data in CIF |  emd_40457_validation.cif.gz emd_40457_validation.cif.gz | 25.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40457 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40457 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40457 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40457 | HTTPS FTP |
-Related structure data
| Related structure data |  8sgjMC  8sgtC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40457.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40457.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: focused CBD map of human cardiac NCX1 in apo inactivated state
| File | emd_40457_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | focused CBD map of human cardiac NCX1 in apo inactivated state | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: focused TM map of human cardiac NCX1 in apo inactivated state
| File | emd_40457_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | focused TM map of human cardiac NCX1 in apo inactivated state | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_40457_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_40457_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human cardiac NCX1 in complex with antibody Fab fragment
| Entire | Name: human cardiac NCX1 in complex with antibody Fab fragment |
|---|---|
| Components |
|
-Supramolecule #1: human cardiac NCX1 in complex with antibody Fab fragment
| Supramolecule | Name: human cardiac NCX1 in complex with antibody Fab fragment type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|
-Supramolecule #2: human cardiac NCX1 in apo inactivated state
| Supramolecule | Name: human cardiac NCX1 in apo inactivated state / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: antibody Fab fragment
| Supramolecule | Name: antibody Fab fragment / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Sodium/calcium exchanger 1
| Macromolecule | Name: Sodium/calcium exchanger 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 109.18107 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MYNMRRLSLS PTFSMGFHLL VTVSLLFSHV DHVIAETEME GEGNETGECT GSYYCKKGVI LPIWEPQDPS FGDKIARATV YFVAMVYMF LGVSIIADRF MSSIEVITSQ EKEITIKKPN GETTKTTVRI WNETVSNLTL MALGSSAPEI LLSVIEVCGH N FTAGDLGP ...String: MYNMRRLSLS PTFSMGFHLL VTVSLLFSHV DHVIAETEME GEGNETGECT GSYYCKKGVI LPIWEPQDPS FGDKIARATV YFVAMVYMF LGVSIIADRF MSSIEVITSQ EKEITIKKPN GETTKTTVRI WNETVSNLTL MALGSSAPEI LLSVIEVCGH N FTAGDLGP STIVGSAAFN MFIIIALCVY VVPDGETRKI KHLRVFFVTA AWSIFAYTWL YIILSVISPG VVEVWEGLLT FF FFPICVV FAWVADRRLL FYKYVYKRYR AGKQRGMIIE HEGDRPSSKT EIEMDGKVVN SHVENFLDGA LVLEVDERDQ DDE EARREM ARILKELKQK HPDKEIEQLI ELANYQVLSQ QQKSRAFYRI QATRLMTGAT ENDPVSKIFF EQGTYQCLEN CGTV ALTII RRGGDLTNTV FVDFRTEDGT ANAGSDYEFT EGTVVFKPGD TQKEIRVGII DDDIFEEDEN FLVHLSNVKV SSEAS EDGI LEANHVSTLA CLGSPSTATV TIFDDDHAGI FTFEEPVTHV SESIGIMEVK VLRTSGARGN VIVPYKTIEG TARGGG EDF EDTCGELEFQ NDEIVKTISV KVIDDEEYEK NKTFFLEIGE PRLVEMSEKK ALLLNELGGF TITGKYLFGQ PVFRKVH AR EHPILSTVIT IADEYDDKQP LTSKEEEERR IAEMGRPILG EHTKLEVIIE ESYEFKSTVD KLIKKTNLAL VVGTNSWR E QFIEAITVSA GEDDDDDECG EEKLPSCFDY VMHFLTVFWK VLFAFVPPTE YWNGWACFIV SILMIGLLTA FIGDLASHF GCTIGLKDSV TAVVFVALGT SVPDTFASKV AATQDQYADA SIGNVTGSNA VNVFLGIGVA WSIAAIYHAA NGEQFKVSPG TLAFSVTLF TIFAFINVGV LLYRRRPEIG GELGGPRTAK LLTSCLFVLL WLLYIFFSSL EAYCHIKGFA AAGGSWSHPQ F EKGGGSGG GSGGSAWSHP QFEK UniProtKB: Sodium/calcium exchanger 1 |
-Macromolecule #2: Fab light chain
| Macromolecule | Name: Fab light chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.740082 KDa |
| Sequence | String: QAVVTQESAL TTSPGETVTL TCRSSTGAVT TSNYANWVQE KPDHLFTGLI GGTNNRVPGV PARFSGSLIG DKAALTITGA QTEDEAIYF CALWYSNHWV FGGGTKLTVL GQPKSSPSVT LFPPSSEELE TNKATLVCTI TDFYPGVVTV DWKVDGTPVT Q GMETTQPS ...String: QAVVTQESAL TTSPGETVTL TCRSSTGAVT TSNYANWVQE KPDHLFTGLI GGTNNRVPGV PARFSGSLIG DKAALTITGA QTEDEAIYF CALWYSNHWV FGGGTKLTVL GQPKSSPSVT LFPPSSEELE TNKATLVCTI TDFYPGVVTV DWKVDGTPVT Q GMETTQPS KQSNNKYMAS SYLTLTARAW ERHSSYSCQV THEG |
-Macromolecule #3: Fab heavy chain
| Macromolecule | Name: Fab heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.435598 KDa |
| Sequence | String: QVQLQQSGAE LARPGASVKL SCKATGYSFT SYWMQWVKQR PGQGMEWIGA IYPGDVTSRY TQKFKGKATL TADKSSSTAF MQLRSLASE DSAVYYCARW SGYYGSSSFD YWGQGTTLTV SSAKTTPPSV YPLAPGCGDT TGSSVTLGCL VKGYFPESVT V TWNSGSLS ...String: QVQLQQSGAE LARPGASVKL SCKATGYSFT SYWMQWVKQR PGQGMEWIGA IYPGDVTSRY TQKFKGKATL TADKSSSTAF MQLRSLASE DSAVYYCARW SGYYGSSSFD YWGQGTTLTV SSAKTTPPSV YPLAPGCGDT TGSSVTLGCL VKGYFPESVT V TWNSGSLS SSVHTFPALL QSGLYTMSSS VTVPSSTWPS QTVTCSVAHP ASSTTVDKKL EPSGPISTIN PCPPCKECHK CP APNLEGG PS |
-Macromolecule #4: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 4 / Number of copies: 6 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #5: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 5 / Number of copies: 3 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 1 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)