+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
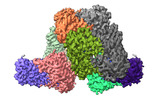
| Title | PRMT1-Decamer | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PRMT1-Decamer / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationGATOR1 complex binding / positive regulation of hemoglobin biosynthetic process / protein-arginine omega-N monomethyltransferase activity / N-methyltransferase activity / peptidyl-arginine methylation / regulation of BMP signaling pathway / regulation of megakaryocyte differentiation / type I protein arginine methyltransferase / protein-arginine omega-N asymmetric methyltransferase activity / histone H4R3 methyltransferase activity ...GATOR1 complex binding / positive regulation of hemoglobin biosynthetic process / protein-arginine omega-N monomethyltransferase activity / N-methyltransferase activity / peptidyl-arginine methylation / regulation of BMP signaling pathway / regulation of megakaryocyte differentiation / type I protein arginine methyltransferase / protein-arginine omega-N asymmetric methyltransferase activity / histone H4R3 methyltransferase activity / protein methyltransferase activity / protein methylation / protein-arginine N-methyltransferase activity / methylosome / cellular response to methionine / S-adenosyl-L-methionine binding / methyl-CpG binding / positive regulation of p38MAPK cascade / cardiac muscle tissue development / negative regulation of JNK cascade / Maturation of nucleoprotein / mitogen-activated protein kinase p38 binding / histone methyltransferase activity / positive regulation of double-strand break repair via homologous recombination / negative regulation of megakaryocyte differentiation / positive regulation of TORC1 signaling / positive regulation of erythrocyte differentiation / TP53 Regulates Transcription of Genes Involved in G2 Cell Cycle Arrest / RNA splicing / methyltransferase activity / positive regulation of translation / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / protein homooligomerization / RMTs methylate histone arginines / neuron projection development / Estrogen-dependent gene expression / in utero embryonic development / cell surface receptor signaling pathway / Extra-nuclear estrogen signaling / viral protein processing / chromatin remodeling / lysosomal membrane / positive regulation of cell population proliferation / DNA damage response / regulation of DNA-templated transcription / negative regulation of apoptotic process / enzyme binding / RNA binding / nucleoplasm / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.56 Å | |||||||||
 Authors Authors | EswarKumar N / Wang CH | |||||||||
| Funding support |  Taiwan, 1 items Taiwan, 1 items
| |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2024 Journal: J Biol Chem / Year: 2024Title: Oligomerization of protein arginine methyltransferase 1 and its functional impact on substrate arginine methylation. Authors: Tran Dang / Nadendla EswarKumar / Sunil Kumar Tripathi / Chunli Yan / Chun-Hsiung Wang / Mengtong Cao / Tanmoy Kumar Paul / Elizabeth Oladoyin Agboluaje / May P Xiong / Ivaylo Ivanov / Meng- ...Authors: Tran Dang / Nadendla EswarKumar / Sunil Kumar Tripathi / Chunli Yan / Chun-Hsiung Wang / Mengtong Cao / Tanmoy Kumar Paul / Elizabeth Oladoyin Agboluaje / May P Xiong / Ivaylo Ivanov / Meng-Chiao Ho / Y George Zheng /   Abstract: Protein arginine methyltransferases (PRMTs) are important posttranslational modifying enzymes in eukaryotic proteins and regulate diverse pathways from gene transcription, RNA splicing, and signal ...Protein arginine methyltransferases (PRMTs) are important posttranslational modifying enzymes in eukaryotic proteins and regulate diverse pathways from gene transcription, RNA splicing, and signal transduction to metabolism. Increasing evidence supports that PRMTs exhibit the capacity to form higher-order oligomeric structures, but the structural basis of PRMT oligomerization and its functional consequence are elusive. Herein, we revealed for the first time different oligomeric structural forms of the predominant arginine methyltransferase PRMT1 using cryo-EM, which included tetramer (dimer of dimers), hexamer (trimer of dimers), octamer (tetramer of dimers), decamer (pentamer of dimers), and also helical filaments. Through a host of biochemical assays, we showed that PRMT1 methyltransferase activity was substantially enhanced as a result of the high-ordered oligomerization. High-ordered oligomerization increased the catalytic turnover and the multimethylation processivity of PRMT1. Presence of a catalytically dead PRMT1 mutant also enhanced the activity of WT PRMT1, pointing out a noncatalytic role of oligomerization. Structural modeling demonstrates that oligomerization enhances substrate retention at the PRMT1 surface through electrostatic force. Our studies offered key insights into PRMT1 oligomerization and established that oligomerization constitutes a novel molecular mechanism that positively regulates the enzymatic activity of PRMTs in biology. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_39814.map.gz emd_39814.map.gz | 203.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-39814-v30.xml emd-39814-v30.xml emd-39814.xml emd-39814.xml | 17.3 KB 17.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_39814.png emd_39814.png | 99.1 KB | ||
| Filedesc metadata |  emd-39814.cif.gz emd-39814.cif.gz | 5.7 KB | ||
| Others |  emd_39814_half_map_1.map.gz emd_39814_half_map_1.map.gz emd_39814_half_map_2.map.gz emd_39814_half_map_2.map.gz | 200 MB 200 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39814 http://ftp.pdbj.org/pub/emdb/structures/EMD-39814 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39814 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39814 | HTTPS FTP |
-Related structure data
| Related structure data |  8z7hMC  8z2zC  8z7oC  9bh4C  9bhdC  9bhgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_39814.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_39814.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_39814_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_39814_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PRMT1-Octamer
| Entire | Name: PRMT1-Octamer |
|---|---|
| Components |
|
-Supramolecule #1: PRMT1-Octamer
| Supramolecule | Name: PRMT1-Octamer / type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Protein arginine N-methyltransferase 1
| Macromolecule | Name: Protein arginine N-methyltransferase 1 / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO / EC number: type I protein arginine methyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.284848 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PNAEDMTSKD YYFDSYAHFG IHEEMLKDEV RTLTYRNSMF HNRHLFKDKV VLDVGSGTGI LCMFAAKAGA RKVIGIECSS ISDYAVKIV KANKLDHVVT IIKGKVEEVE LPVEKVDIII SEWMGYCLFY ESMLNTVLYA RDKWLAPDGL IFPDRATLYV T AIEDRQYK ...String: PNAEDMTSKD YYFDSYAHFG IHEEMLKDEV RTLTYRNSMF HNRHLFKDKV VLDVGSGTGI LCMFAAKAGA RKVIGIECSS ISDYAVKIV KANKLDHVVT IIKGKVEEVE LPVEKVDIII SEWMGYCLFY ESMLNTVLYA RDKWLAPDGL IFPDRATLYV T AIEDRQYK DYKIHWWENV YGFDMSCIKD VAIKEPLVDV VDPKQLVTNA CLIKEVDIYT VKVEDLTFTS PFCLQVKRND YV HALVAYF NIEFTRCHKR TGFSTSPESP YTHWKQTVFY MEDYLTVKTG EEIFGTIGMR PNAKNNRDLD FTIDLDFKGQ LCE LSCSTD YRMR UniProtKB: Protein arginine N-methyltransferase 1 |
-Macromolecule #2: S-ADENOSYL-L-HOMOCYSTEINE
| Macromolecule | Name: S-ADENOSYL-L-HOMOCYSTEINE / type: ligand / ID: 2 / Number of copies: 8 / Formula: SAH |
|---|---|
| Molecular weight | Theoretical: 384.411 Da |
| Chemical component information |  ChemComp-SAH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 23.34 Å Applied symmetry - Helical parameters - Δ&Phi: 103.59 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 3.56 Å / Resolution method: FSC 3 SIGMA CUT-OFF / Number images used: 541164 |
|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
| Startup model | Type of model: NONE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































