+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3911 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bdellovibrio bacteriovorus bacterial flagellar motor | |||||||||
 Map data Map data | Bdellovibrio bacteriovorus bacterial flagellar motor | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Bdellovibrio bacteriovorus (bacteria) Bdellovibrio bacteriovorus (bacteria) | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 82.0 Å | |||||||||
 Authors Authors | Chaban B / Coleman I / Beeby M | |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2018 Journal: Sci Rep / Year: 2018Title: Evolution of higher torque in Campylobacter-type bacterial flagellar motors. Authors: Bonnie Chaban / Izaak Coleman / Morgan Beeby /   Abstract: Understanding the evolution of molecular machines underpins our understanding of the development of life on earth. A well-studied case are bacterial flagellar motors that spin helical propellers for ...Understanding the evolution of molecular machines underpins our understanding of the development of life on earth. A well-studied case are bacterial flagellar motors that spin helical propellers for bacterial motility. Diverse motors produce different torques, but how this diversity evolved remains unknown. To gain insights into evolution of the high-torque ε-proteobacterial motor exemplified by the Campylobacter jejuni motor, we inferred ancestral states by combining phylogenetics, electron cryotomography, and motility assays to characterize motors from Wolinella succinogenes, Arcobacter butzleri and Bdellovibrio bacteriovorus. Observation of ~12 stator complexes in many proteobacteria, yet ~17 in ε-proteobacteria suggest a "quantum leap" evolutionary event. Campylobacter-type motors have high stator occupancy in wider rings of additional stator complexes that are scaffolded by large proteinaceous periplasmic rings. We propose a model for motor evolution wherein independent inner- and outer-membrane structures fused to form a scaffold for additional stator complexes. Significantly, inner- and outer-membrane associated structures have evolved independently multiple times, suggesting that evolution of such structures is facile and poised the ε-proteobacteria to fuse them to form the high-torque Campylobacter-type motor. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3911.map.gz emd_3911.map.gz | 22.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3911-v30.xml emd-3911-v30.xml emd-3911.xml emd-3911.xml | 7.3 KB 7.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3911.png emd_3911.png | 53 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3911 http://ftp.pdbj.org/pub/emdb/structures/EMD-3911 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3911 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3911 | HTTPS FTP |
-Validation report
| Summary document |  emd_3911_validation.pdf.gz emd_3911_validation.pdf.gz | 198.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3911_full_validation.pdf.gz emd_3911_full_validation.pdf.gz | 197.2 KB | Display | |
| Data in XML |  emd_3911_validation.xml.gz emd_3911_validation.xml.gz | 6.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3911 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3911 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3911 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3911 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3911.map.gz / Format: CCP4 / Size: 24.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3911.map.gz / Format: CCP4 / Size: 24.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Bdellovibrio bacteriovorus bacterial flagellar motor | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 8.122 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
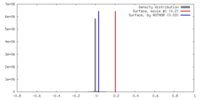
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bdellovibrio bacteriovorus flagellar motor
| Entire | Name: Bdellovibrio bacteriovorus flagellar motor |
|---|---|
| Components |
|
-Supramolecule #1: Bdellovibrio bacteriovorus flagellar motor
| Supramolecule | Name: Bdellovibrio bacteriovorus flagellar motor / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Bdellovibrio bacteriovorus (bacteria) Bdellovibrio bacteriovorus (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: Quantifoil R2/2 |
| Vitrification | Cryogen name: ETHANE-PROPANE |
| Details | In situ |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 20 |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 82.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: PEET / Number subtomograms used: 478 |
|---|---|
| Extraction | Number tomograms: 478 / Number images used: 478 |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)