+ Open data
Open data
- Basic information
Basic information
| Entry | 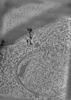 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SPACEtomo example tomograms of Baker's Yeast | |||||||||
 Map data Map data | SPACEtomo example tomogram showing a mitochondrium | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cryoET / tomogram / CYTOSOLIC PROTEIN | |||||||||
| Biological species |  | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Eisenstein F / Danev R | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2024 Journal: Nat Methods / Year: 2024Title: Smart parallel automated cryo-electron tomography. Authors: Fabian Eisenstein / Yoshiyuki Fukuda / Radostin Danev /   Abstract: In situ cryo-electron tomography enables investigation of macromolecules in their native cellular environment. Samples have become more readily available owing to recent software and hardware ...In situ cryo-electron tomography enables investigation of macromolecules in their native cellular environment. Samples have become more readily available owing to recent software and hardware advancements. Data collection, however, still requires an experienced operator and appreciable microscope time to carefully select targets for high-throughput tilt series acquisition. Here, we developed smart parallel automated cryo-electron tomography (SPACEtomo), a workflow using machine learning approaches to fully automate the entire cryo-electron tomography process, including lamella detection, biological feature segmentation, target selection and parallel tilt series acquisition, all without the need for human intervention. This degree of automation will be essential for obtaining statistically relevant datasets and high-resolution structures of macromolecules in their native context. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38295.map.gz emd_38295.map.gz | 1.5 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38295-v30.xml emd-38295-v30.xml emd-38295.xml emd-38295.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_38295.png emd_38295.png | 167.5 KB | ||
| Filedesc metadata |  emd-38295.cif.gz emd-38295.cif.gz | 3.8 KB | ||
| Others |  emd_38295_additional_1.map.gz emd_38295_additional_1.map.gz | 2.5 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38295 http://ftp.pdbj.org/pub/emdb/structures/EMD-38295 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38295 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38295 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_38295.map.gz / Format: CCP4 / Size: 1.6 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38295.map.gz / Format: CCP4 / Size: 1.6 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SPACEtomo example tomogram showing a mitochondrium | ||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 8.36 Å | ||||||||||||||||||||||||||||||||
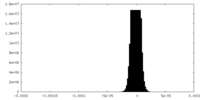
| Density |
| ||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: SPACEtomo example tomogram showing the nuclear envelope
| File | emd_38295_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SPACEtomo example tomogram showing the nuclear envelope | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Baker's Yeast
| Entire | Name:  |
|---|---|
| Components |
|
-Supramolecule #1: Baker's Yeast
| Supramolecule | Name: Baker's Yeast / type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
| Cryo protectant | 15 % dextran and 2 % glucose |
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 / Focused ion beam - Current: 0.03 / Focused ion beam - Duration: 60 / Focused ion beam - Temperature: 78 K / Focused ion beam - Initial thickness: 1000 / Focused ion beam - Final thickness: 200 Focused ion beam - Details: The value given for _em_focused_ion_beam.instrument is Aquilos 2. This is not in a list of allowed values {'OTHER', 'DB235'} so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 8 / Average exposure time: 1.0 sec. / Average electron dose: 4.35 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 5.0 µm / Nominal magnification: 42000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: BACK PROJECTION / Details: Tomogram was denoised using cryo-CARE / Number images used: 40 |
|---|
 Movie
Movie Controller
Controller
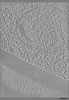




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
























