[English] 日本語
 Yorodumi
Yorodumi- EMDB-37829: Cryo-EM structure of the IS621 recombinase in complex with bridge... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the IS621 recombinase in complex with bridge RNA, donor DNA, and target DNA in the post-strand exchange state (Holliday junction intermediate) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Holliday junction / RNA dependent recombinase / RECOMBINATION-RNA-DNA COMPLEX | |||||||||
| Function / homology | Transposase, IS111A/IS1328/IS1533, N-terminal / : / Transposase / Transposase, IS116/IS110/IS902 / Transposase IS116/IS110/IS902 family / transposase activity / DNA transposition / DNA binding / IS621 transposase Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
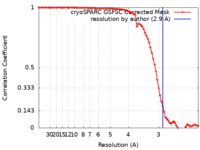
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Hiraizumi M / Yamashita K / Nishimasu H | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Structural mechanism of bridge RNA-guided recombination. Authors: Masahiro Hiraizumi / Nicholas T Perry / Matthew G Durrant / Teppei Soma / Naoto Nagahata / Sae Okazaki / Januka S Athukoralage / Yukari Isayama / James J Pai / April Pawluk / Silvana ...Authors: Masahiro Hiraizumi / Nicholas T Perry / Matthew G Durrant / Teppei Soma / Naoto Nagahata / Sae Okazaki / Januka S Athukoralage / Yukari Isayama / James J Pai / April Pawluk / Silvana Konermann / Keitaro Yamashita / Patrick D Hsu / Hiroshi Nishimasu /   Abstract: Insertion sequence (IS) elements are the simplest autonomous transposable elements found in prokaryotic genomes. We recently discovered that IS110 family elements encode a recombinase and a non- ...Insertion sequence (IS) elements are the simplest autonomous transposable elements found in prokaryotic genomes. We recently discovered that IS110 family elements encode a recombinase and a non-coding bridge RNA (bRNA) that confers modular specificity for target DNA and donor DNA through two programmable loops. Here we report the cryo-electron microscopy structures of the IS110 recombinase in complex with its bRNA, target DNA and donor DNA in three different stages of the recombination reaction cycle. The IS110 synaptic complex comprises two recombinase dimers, one of which houses the target-binding loop of the bRNA and binds to target DNA, whereas the other coordinates the bRNA donor-binding loop and donor DNA. We uncovered the formation of a composite RuvC-Tnp active site that spans the two dimers, positioning the catalytic serine residues adjacent to the recombination sites in both target and donor DNA. A comparison of the three structures revealed that (1) the top strands of target and donor DNA are cleaved at the composite active sites to form covalent 5'-phosphoserine intermediates, (2) the cleaved DNA strands are exchanged and religated to create a Holliday junction intermediate, and (3) this intermediate is subsequently resolved by cleavage of the bottom strands. Overall, this study reveals the mechanism by which a bispecific RNA confers target and donor DNA specificity to IS110 recombinases for programmable DNA recombination. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37829.map.gz emd_37829.map.gz | 26.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37829-v30.xml emd-37829-v30.xml emd-37829.xml emd-37829.xml | 19.7 KB 19.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37829_fsc.xml emd_37829_fsc.xml | 7.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_37829.png emd_37829.png | 131.2 KB | ||
| Masks |  emd_37829_msk_1.map emd_37829_msk_1.map | 52.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-37829.cif.gz emd-37829.cif.gz | 6.4 KB | ||
| Others |  emd_37829_half_map_1.map.gz emd_37829_half_map_1.map.gz emd_37829_half_map_2.map.gz emd_37829_half_map_2.map.gz | 49 MB 49 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37829 http://ftp.pdbj.org/pub/emdb/structures/EMD-37829 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37829 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37829 | HTTPS FTP |
-Related structure data
| Related structure data |  8wt8MC  8wt6C  8wt7C  8wt9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_37829.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37829.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.10667 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_37829_msk_1.map emd_37829_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_37829_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_37829_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : The IS621 recombinase in complex with bridge RNA, donor DNA, and ...
| Entire | Name: The IS621 recombinase in complex with bridge RNA, donor DNA, and target DNA in the post-strand exchange state (Holliday junction intermediate) |
|---|---|
| Components |
|
-Supramolecule #1: The IS621 recombinase in complex with bridge RNA, donor DNA, and ...
| Supramolecule | Name: The IS621 recombinase in complex with bridge RNA, donor DNA, and target DNA in the post-strand exchange state (Holliday junction intermediate) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: IS621 transposase
| Macromolecule | Name: IS621 transposase / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.735355 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPMEHELHYI GIDTAKEKLD VDVLRPDGRH RTKKFANTTK GHDELVSWLK GHKIDHAHIC IEATGTYMEP VAECLYDAGY IVSVINPAL GKAFAQSEGL RNKTDTVDAR MLAEFCRQKR PAAWEAPHPL ERALRALVVR HQALTDMHTQ ELNRTETARE V QRPSIDAH ...String: GPMEHELHYI GIDTAKEKLD VDVLRPDGRH RTKKFANTTK GHDELVSWLK GHKIDHAHIC IEATGTYMEP VAECLYDAGY IVSVINPAL GKAFAQSEGL RNKTDTVDAR MLAEFCRQKR PAAWEAPHPL ERALRALVVR HQALTDMHTQ ELNRTETARE V QRPSIDAH LLWLEAELKR LEKQIKDLTD DDPDMKHRRK LLESIPGIGE KTSAVLLAYI GLKDRFAHAR QFAAFAGLTP RR YESGSSV RGASRMSKAG HVSLRRALYM PAMVATSKTE WGRAFRDRLA ANGKKGKVIL GAMMRKLAQV AYGVLKSGVP FDA SRHNPV AA UniProtKB: IS621 transposase |
-Macromolecule #2: bridge RNA
| Macromolecule | Name: bridge RNA / type: rna / ID: 2 / Number of copies: 2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 57.668918 KDa |
| Sequence | String: GGGAGUGCAG AGAAAAUCGG CCAGUUUUCU CUGCCUGCAG UCCGCAUGCC GUAUCGGGCC UUGGGUUCUA ACCUGUUCUG UAGAUUUAU GCAGCGGACU GCCUUUCUCC CAAAGUGAUA AACCGGACAG UAUCAUGGAC CGGUUUUCCC GGUAAUCCGU A UUUACAAG GCUGGUUUCA CU |
-Macromolecule #3: target DNA-donor DNA
| Macromolecule | Name: target DNA-donor DNA / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.003622 KDa |
| Sequence | String: (DG)(DC)(DC)(DG)(DG)(DG)(DT)(DA)(DA)(DT) (DA)(DC)(DC)(DA)(DC)(DC)(DA)(DA)(DG)(DC) (DC)(DG)(DC)(DC)(DT)(DT)(DG)(DT)(DA) (DT)(DT)(DA)(DT)(DC)(DC)(DC)(DT)(DC)(DC) (DA) (DG)(DT)(DG)(DC)(DA)(DG)(DA)(DG) (DA) |
-Macromolecule #4: target DNA
| Macromolecule | Name: target DNA / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.703487 KDa |
| Sequence | String: (DC)(DG)(DA)(DG)(DC)(DT)(DC)(DA)(DT)(DC) (DT)(DG)(DT)(DA)(DG)(DG)(DC)(DC)(DC)(DG) (DA)(DT)(DG)(DG)(DT)(DG)(DG)(DT)(DA) (DT)(DT)(DA)(DC)(DC)(DC)(DG)(DG)(DC) |
-Macromolecule #5: donor DNA-target DNA
| Macromolecule | Name: donor DNA-target DNA / type: dna / ID: 5 Details: The linkage between 20I and 21I has been cleaved by IS621. Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 10.163561 KDa |
| Sequence | String: (DT)(DG)(DC)(DA)(DG)(DG)(DC)(DC)(DA)(DT) (DA)(DA)(DG)(DT)(DC)(DA)(DA)(DT)(DC)(DT) (DA)(DC)(DA)(DG)(DA)(DT)(DG)(DA)(DG) (DC)(DT)(DC)(DG) |
-Macromolecule #6: donor DNA
| Macromolecule | Name: donor DNA / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.587737 KDa |
| Sequence | String: (DT)(DC)(DT)(DC)(DT)(DG)(DC)(DA)(DC)(DT) (DG)(DG)(DA)(DG)(DG)(DG)(DA)(DT)(DA)(DA) (DT)(DA)(DC)(DA)(DA)(DG)(DA)(DT)(DA) (DC)(DT)(DG)(DT)(DT)(DA)(DT)(DG)(DG)(DC) (DC) (DT)(DG)(DC)(DA) |
-Macromolecule #7: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 7 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 62.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)