+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of OAT4 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SLC / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationOrganic anion transport by SLC22 transporters / : / prostaglandin transport / prostaglandin transmembrane transporter activity / urate metabolic process / solute:inorganic anion antiporter activity / : / : / monoatomic ion transport / basal plasma membrane ...Organic anion transport by SLC22 transporters / : / prostaglandin transport / prostaglandin transmembrane transporter activity / urate metabolic process / solute:inorganic anion antiporter activity / : / : / monoatomic ion transport / basal plasma membrane / apical plasma membrane / external side of plasma membrane / extracellular exosome / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Qian HW / He JJ | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2024 Journal: Cell Rep / Year: 2024Title: Structural basis for the transport and substrate selection of human urate transporter 1. Authors: Jingjing He / Guoyun Liu / Fang Kong / Qiulong Tan / Zhenzhou Wang / Meng Yang / Yonglin He / Xiaoxiao Jia / Chuangye Yan / Chao Wang / Hongwu Qian /  Abstract: High serum urate levels are the major risk factor for gout. URAT1, the primary transporter for urate absorption in the kidneys, is well known as an anti-hyperuricemia drug target. However, the ...High serum urate levels are the major risk factor for gout. URAT1, the primary transporter for urate absorption in the kidneys, is well known as an anti-hyperuricemia drug target. However, the clinical application of URAT1-targeted drugs is limited because of their low specificity and severe side effects. The lack of structural information impedes elucidation of the transport mechanism and the development of new drugs. Here, we present the cryoelectron microscopy (cryo-EM) structures of human URAT1(R477S), its complex with urate, and its closely related homolog OAT4. URAT1(R477S) and OAT4 exhibit major facilitator superfamily (MFS) folds with outward- and inward-open conformations, respectively. Structural comparison reveals a 30° rotation between the N-terminal and C-terminal domains, supporting an alternating access mechanism. A conserved arginine (OAT4-Arg473/URAT1-Arg477) is found to be essential for chloride-mediated inhibition. The URAT1(R477S)-urate complex reveals the specificity of urate recognition. Taken together, our study promotes our understanding of the transport mechanism and substrate selection of URAT1. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37580.map.gz emd_37580.map.gz | 20.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37580-v30.xml emd-37580-v30.xml emd-37580.xml emd-37580.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_37580.png emd_37580.png | 38.6 KB | ||
| Filedesc metadata |  emd-37580.cif.gz emd-37580.cif.gz | 5.8 KB | ||
| Others |  emd_37580_half_map_1.map.gz emd_37580_half_map_1.map.gz emd_37580_half_map_2.map.gz emd_37580_half_map_2.map.gz | 20.6 MB 20.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37580 http://ftp.pdbj.org/pub/emdb/structures/EMD-37580 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37580 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37580 | HTTPS FTP |
-Related structure data
| Related structure data |  8wjhMC  8wjgC  8wjqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37580.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37580.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_37580_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
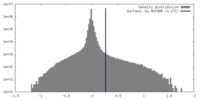
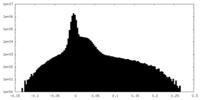
| Projections & Slices |
| ||||||||||||
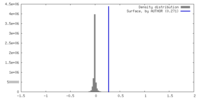
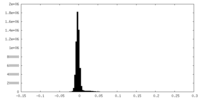
| Density Histograms |
-Half map: #1
| File | emd_37580_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of OAT4
| Entire | Name: Cryo-EM structure of OAT4 |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of OAT4
| Supramolecule | Name: Cryo-EM structure of OAT4 / type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Solute carrier family 22 member 11
| Macromolecule | Name: Solute carrier family 22 member 11 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.506453 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAFSKLLEQA GGVGLFQTLQ VLTFILPCLM IPSQMLLENF SAAIPGHRCW THMLDNGSAV STNMTPKALL TISIPPGPNQ GPHQCRRFR QPQWQLLDPN ATATSWSEAD TEPCVDGWVY DRSVFTSTIV AKWDLVCSSQ GLKPLSQSIF MSGILVGSFI W GLLSYRFG ...String: MAFSKLLEQA GGVGLFQTLQ VLTFILPCLM IPSQMLLENF SAAIPGHRCW THMLDNGSAV STNMTPKALL TISIPPGPNQ GPHQCRRFR QPQWQLLDPN ATATSWSEAD TEPCVDGWVY DRSVFTSTIV AKWDLVCSSQ GLKPLSQSIF MSGILVGSFI W GLLSYRFG RKPMLSWCCL QLAVAGTSTI FAPTFVIYCG LRFVAAFGMA GIFLSSLTLM VEWTTTSRRA VTMTVVGCAF SA GQAALGG LAFALRDWRT LQLAASVPFF AISLISWWLP ESARWLIIKG KPDQALQELR KVARINGHKE AKNLTIEVLM SSV KEEVAS AKEPRSVLDL FCVPVLRWRS CAMLVVNFSL LISYYGLVFD LQSLGRDIFL LQALFGAVDF LGRATTALLL SFLG RRTIQ AGSQAMAGLA ILANMLVPQD LQTLRVVFAV LGKGCFGISL TCLTIYKAEL FPTPVRMTAD GILHTVGRLG AMMGP LILM SRQALPLLPP LLYGVISIAS SLVVLFFLPE TQG UniProtKB: Solute carrier family 22 member 11 |
-Macromolecule #2: CHLORIDE ION
| Macromolecule | Name: CHLORIDE ION / type: ligand / ID: 2 / Number of copies: 1 / Formula: CL |
|---|---|
| Molecular weight | Theoretical: 35.453 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.7 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)