+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CXCR3-DNGi complex activated by CXCL10 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | G protein coupled receptor / chemokine receptor / CXCR3 / CXCL10 / complex / SIGNALING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of T cell chemotaxis / negative regulation of myoblast fusion / Oplophorus-luciferin 2-monooxygenase / Oplophorus-luciferin 2-monooxygenase activity / CXCR3 chemokine receptor binding / regulation of leukocyte migration / chemokine binding / C-X-C chemokine binding / regulation of endothelial tube morphogenesis / cellular response to interleukin-17 ...regulation of T cell chemotaxis / negative regulation of myoblast fusion / Oplophorus-luciferin 2-monooxygenase / Oplophorus-luciferin 2-monooxygenase activity / CXCR3 chemokine receptor binding / regulation of leukocyte migration / chemokine binding / C-X-C chemokine binding / regulation of endothelial tube morphogenesis / cellular response to interleukin-17 / chemokine receptor activity / CXCR chemokine receptor binding / C-X-C chemokine receptor activity / cAMP-dependent protein kinase regulator activity / response to auditory stimulus / negative regulation of myoblast differentiation / positive regulation of chemotaxis / T cell chemotaxis / C-C chemokine receptor activity / C-C chemokine binding / negative regulation of execution phase of apoptosis / positive regulation of monocyte chemotaxis / endothelial cell activation / response to vitamin D / chemokine activity / Chemokine receptors bind chemokines / muscle organ development / blood circulation / negative regulation of endothelial cell proliferation / chemoattractant activity / Interleukin-10 signaling / positive regulation of execution phase of apoptosis / positive regulation of T cell migration / regulation of cell adhesion / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration / positive regulation of relaxation of smooth muscle / neutrophil chemotaxis / Adenylate cyclase inhibitory pathway / D2 dopamine receptor binding / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / antiviral innate immune response / negative regulation of angiogenesis / cellular response to forskolin / bioluminescence / positive regulation of release of sequestered calcium ion into cytosol / regulation of mitotic spindle organization / chemokine-mediated signaling pathway / response to gamma radiation / cell chemotaxis / Regulation of insulin secretion / neuropeptide signaling pathway / calcium-mediated signaling / response to prostaglandin E / positive regulation of cholesterol biosynthetic process / cellular response to virus / electron transport chain / negative regulation of insulin secretion / G protein-coupled receptor binding / response to peptide hormone / G-protein beta/gamma-subunit complex binding / centriolar satellite / chemotaxis / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / positive regulation of angiogenesis / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G beta:gamma signalling through BTK / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / GDP binding / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / cell-cell signaling / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / regulation of cell population proliferation / heparin binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Oplophorus gracilirostris (arthropod) Oplophorus gracilirostris (arthropod) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Jiao HZ / Hu HL | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2023 Journal: Cell Discov / Year: 2023Title: Structure basis for the modulation of CXC chemokine receptor 3 by antagonist AMG487. Authors: Haizhan Jiao / Bin Pang / Ying-Chih Chiang / Qiang Chen / Qi Pan / Ruobing Ren / Hongli Hu /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36842.map.gz emd_36842.map.gz | 66.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36842-v30.xml emd-36842-v30.xml emd-36842.xml emd-36842.xml | 29.1 KB 29.1 KB | Display Display |  EMDB header EMDB header |
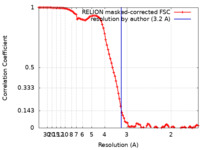
| FSC (resolution estimation) |  emd_36842_fsc.xml emd_36842_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_36842.png emd_36842.png | 105.6 KB | ||
| Filedesc metadata |  emd-36842.cif.gz emd-36842.cif.gz | 8 KB | ||
| Others |  emd_36842_additional_1.map.gz emd_36842_additional_1.map.gz emd_36842_additional_2.map.gz emd_36842_additional_2.map.gz emd_36842_half_map_1.map.gz emd_36842_half_map_1.map.gz emd_36842_half_map_2.map.gz emd_36842_half_map_2.map.gz | 41.4 MB 65.1 MB 65.4 MB 65.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36842 http://ftp.pdbj.org/pub/emdb/structures/EMD-36842 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36842 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36842 | HTTPS FTP |
-Related structure data
| Related structure data |  8k2xMC  8k2wC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36842.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36842.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: local scale map
| File | emd_36842_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | local scale map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: unsharpen map
| File | emd_36842_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpen map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_36842_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36842_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CXCR3-CXCL10-DNGi-scFv16
| Entire | Name: CXCR3-CXCL10-DNGi-scFv16 |
|---|---|
| Components |
|
-Supramolecule #1: CXCR3-CXCL10-DNGi-scFv16
| Supramolecule | Name: CXCR3-CXCL10-DNGi-scFv16 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.445059 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGAQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHESM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCS TDTKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.005895 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: HHHHHHHHHH LEVLFQGPGS SGSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSR LLVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE L AGHTGYLS ...String: HHHHHHHHHH LEVLFQGPGS SGSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSR LLVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE L AGHTGYLS CCRFLDDNQI VTSSGDTTCA LWDIETGQQT TTFTGHTGDV MSLSLAPDTR LFVSGACDAS AKLWDVREGM CR QTFTGHE SDINAICFFP NGNAFATGSD DATCRLFDLR ADQELMTYSH DNIICGITSV SFSKSGRLLL AGYDDFNCNV WDA LKADRA GVLAGHDNRV SCLGVTDDGM AVATGSWDSF LKIWNGASGA SGASGASVSG WRLFKKIS UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Soluble cytochrome b562,C-X-C chemokine receptor type 3,Oplophoru...
| Macromolecule | Name: Soluble cytochrome b562,C-X-C chemokine receptor type 3,Oplophorus-luciferin 2-monooxygenase catalytic subunit type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO / EC number: Oplophorus-luciferin 2-monooxygenase |
|---|---|
| Source (natural) | Organism:  Oplophorus gracilirostris (arthropod) Oplophorus gracilirostris (arthropod) |
| Molecular weight | Theoretical: 72.918008 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDKGSADLE DNWETLNDNL KVIEKADNAA QVKDALTKMR AAALDAQKAT PPKLEDKSPD SPEMKDFRH GFDILVGQID DALKLANEGK VKEAQAAAEQ LKTTRNAYIQ KYLLVPRGSM VLEVSDHQVL NDAEVAALLE N FSSSYDYG ...String: MKTIIALSYI FCLVFADYKD DDDKGSADLE DNWETLNDNL KVIEKADNAA QVKDALTKMR AAALDAQKAT PPKLEDKSPD SPEMKDFRH GFDILVGQID DALKLANEGK VKEAQAAAEQ LKTTRNAYIQ KYLLVPRGSM VLEVSDHQVL NDAEVAALLE N FSSSYDYG ENESDSCCTS PPCPQDFSLN FDRAFLPALY SLLFLLGLLG NGAVAAVLLS RRTALSSTDT FLLHLAVADT LL VLTLPLW AVDAAVQWVF GSGLCKVAGA LFNINFYAGA LLLACISFDR YLNIVHATQL YRRGPPARVT LTCLAVWGLC LLF ALPDFI FLSAHHDERL NATHCQYNFP QVGRTALRVL QLVAGFLLPL LVMAYCYAHI LAVLLVSRGQ RRLRAMRLVV VVVV AFALC WTPYHLVVLV DILMDLGALA RNCGRESRVD VAKSVTSGLG YMHCCLNPLL YAFVGVKFRE RMWMLLLRLG CPNQR GLQR QPSSSRRDSS WSETSVFTLE DFVGDWEQTA AYNLDQVLEQ GGVSSLLQNL AVSVTPIQRI VRSGENALKI DIHVII PYE GLSADQMAQI EEVFKVVYPV DDHHFKVILP YGTLVIDGVT PNMLNYFGRP YEGIAVFDGK KITVTGTLWN GNKIIDE RL ITPDGSMLFR VTINS UniProtKB: Soluble cytochrome b562, C-X-C chemokine receptor type 3, Oplophorus-luciferin 2-monooxygenase catalytic subunit |
-Macromolecule #5: single-chain variable fragment scFv16
| Macromolecule | Name: single-chain variable fragment scFv16 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Oplophorus gracilirostris (arthropod) Oplophorus gracilirostris (arthropod) |
| Molecular weight | Theoretical: 27.784896 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAAHHHHHH HH |
-Macromolecule #6: C-X-C motif chemokine 10
| Macromolecule | Name: C-X-C motif chemokine 10 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.898089 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNQTAILICC LIFLTLSGIQ GVPLSRTVRC TCISISNQPV NPRSLEKLEI IPASQFCPRV EIIATMKKKG EKRCLNPESK AIKNLLKAV SKERSKRSP UniProtKB: C-X-C motif chemokine 10 |
-Macromolecule #7: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 7 / Number of copies: 1 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5.0 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 53.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)