[English] 日本語
 Yorodumi
Yorodumi- EMDB-36486: Cryo-EM structure of succinate receptor bound to succinate acid c... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of succinate receptor bound to succinate acid coupling MiniGsq | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of angiotensin metabolic process / renin secretion into blood stream / positive regulation of chemotaxis / Class A/1 (Rhodopsin-like receptors) / macrophage activation involved in immune response / adenylate cyclase-activating G protein-coupled bile acid receptor signaling pathway / adenylate cyclase-activating serotonin receptor signaling pathway / regulation of skeletal muscle contraction / PKA activation in glucagon signalling / hair follicle placode formation ...regulation of angiotensin metabolic process / renin secretion into blood stream / positive regulation of chemotaxis / Class A/1 (Rhodopsin-like receptors) / macrophage activation involved in immune response / adenylate cyclase-activating G protein-coupled bile acid receptor signaling pathway / adenylate cyclase-activating serotonin receptor signaling pathway / regulation of skeletal muscle contraction / PKA activation in glucagon signalling / hair follicle placode formation / developmental growth / intracellular transport / D1 dopamine receptor binding / vascular endothelial cell response to laminar fluid shear stress / renal water homeostasis / Hedgehog 'off' state / activation of adenylate cyclase activity / cellular response to acidic pH / adenylate cyclase-activating adrenergic receptor signaling pathway / energy homeostasis / cellular response to glucagon stimulus / intracellular glucose homeostasis / positive regulation of insulin secretion involved in cellular response to glucose stimulus / adenylate cyclase activator activity / trans-Golgi network membrane / negative regulation of inflammatory response to antigenic stimulus / response to prostaglandin E / electron transport chain / bone development / response to calcium ion / platelet aggregation / G protein-coupled receptor activity / cognition / G-protein beta/gamma-subunit complex binding / Olfactory Signaling Pathway / Activation of the phototransduction cascade / positive regulation of insulin secretion / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / positive regulation of inflammatory response / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / sensory perception of smell / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / glucose homeostasis / G alpha (12/13) signalling events / heterotrimeric G-protein complex / positive regulation of cold-induced thermogenesis / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / signaling receptor activity / retina development in camera-type eye / GTPase binding / G protein activity / fibroblast proliferation / Ca2+ pathway / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Ras protein signal transduction / periplasmic space / electron transfer activity / Extra-nuclear estrogen signaling / cell population proliferation / iron ion binding / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase activity / heme binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Oplophorus gracilirostris (arthropod) Oplophorus gracilirostris (arthropod) | |||||||||
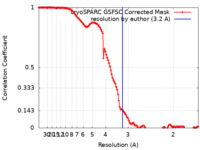
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Wang TX / Tang WQ / Li FH / Wang JY | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2024 Journal: Cell Res / Year: 2024Title: Molecular activation and G protein coupling selectivity of human succinate receptor SUCR1. Authors: Tianxin Wang / Wenqin Tang / Xiaolei Zhu / Zhenyu Lv / Jiayan Chen / Yongze Li / Xiaoyu Sun / Haoyu Lv / Quanchang Gu / Fahui Li / Jiangyun Wang /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36486.map.gz emd_36486.map.gz | 167.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36486-v30.xml emd-36486-v30.xml emd-36486.xml emd-36486.xml | 17.9 KB 17.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36486_fsc.xml emd_36486_fsc.xml | 13.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_36486.png emd_36486.png | 29.3 KB | ||
| Filedesc metadata |  emd-36486.cif.gz emd-36486.cif.gz | 6.4 KB | ||
| Others |  emd_36486_half_map_1.map.gz emd_36486_half_map_1.map.gz emd_36486_half_map_2.map.gz emd_36486_half_map_2.map.gz | 164.9 MB 164.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36486 http://ftp.pdbj.org/pub/emdb/structures/EMD-36486 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36486 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36486 | HTTPS FTP |
-Related structure data
| Related structure data |  8jppMC  8jpnC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36486.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36486.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36486_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36486_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GPCR complex
| Entire | Name: GPCR complex |
|---|---|
| Components |
|
-Supramolecule #1: GPCR complex
| Supramolecule | Name: GPCR complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 29.245139 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGADN SGKSTIVKQM RILHGGSGGS GGTSGIFETK FQVDKVNFH MFDVGGQRDE RRKWIQCFND VTAIIFVVDS SDYNRLQEAL NLFKSIWNNR WLRTISVILF LNKQDLLAEK V LAGKSKIE ...String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGADN SGKSTIVKQM RILHGGSGGS GGTSGIFETK FQVDKVNFH MFDVGGQRDE RRKWIQCFND VTAIIFVVDS SDYNRLQEAL NLFKSIWNNR WLRTISVILF LNKQDLLAEK V LAGKSKIE DYFPEFARYT TPEDATPEPG EDPRVTRAKY FIRDEFLRIS TASGDGRHYC YPHFTCAVDT ENARRIFNDC KD IILQMNL REYNLV UniProtKB: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.055867 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MHHHHHHGSL LQSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSRL LVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE LAGHTGYLSC C RFLDDNQI ...String: MHHHHHHGSL LQSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSRL LVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE LAGHTGYLSC C RFLDDNQI VTSSGDTTCA LWDIETGQQT TTFTGHTGDV MSLSLAPDTR LFVSGACDAS AKLWDVREGM CRQTFTGHES DI NAICFFP NGNAFATGSD DATCRLFDLR ADQELMTYSH DNIICGITSV SFSKSGRLLL AGYDDFNCNV WDALKADRAG VLA GHDNRV SCLGVTDDGM AVATGSWDSF LKIWNGSSGG GGSGGGGSSG VSGWRLFKKI S UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Soluble cytochrome b562,Succinate receptor 1,Oplophorus-luciferin...
| Macromolecule | Name: Soluble cytochrome b562,Succinate receptor 1,Oplophorus-luciferin 2-monooxygenase catalytic subunit type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Oplophorus gracilirostris (arthropod) Oplophorus gracilirostris (arthropod) |
| Molecular weight | Theoretical: 74.993492 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDAGRAHHH HHHHHHHENL YFQSGAPADL EDNWETLNDN LKVIEKADNA AQVKDALTKM RAAALDAQK ATPPKLEDKS PDSPEMKDFR HGFDILVGQI DDALKLANEG KVKEAQAAAE QLKTTRNAYI QKYLLEVLFQ G PEFMLGIM ...String: MKTIIALSYI FCLVFADYKD DDDAGRAHHH HHHHHHHENL YFQSGAPADL EDNWETLNDN LKVIEKADNA AQVKDALTKM RAAALDAQK ATPPKLEDKS PDSPEMKDFR HGFDILVGQI DDALKLANEG KVKEAQAAAE QLKTTRNAYI QKYLLEVLFQ G PEFMLGIM AWNATCKNWL AAEAALEKYY LSIFYGIEFV VGVLGNTIVV YGYIFSLKNW NSSNIYLFNL SVSDLAFLCT LP MLIRSYA NGNWIYGDVL CISNRYVLHA NLYTSILFLT FISIDRYLII KYPFREHLLQ KKEFAILISL AIWVLVTLEL LPI LPLINP VITDNGTTCN DFASSGDPNY NLIYSMCLTL LGFLIPLFVM CFFYYKIALF LKQRNRQVAT ALPLEKPLNL VIMA VVIFS VLFTPYHVMR NVRIASRLGS WKQYQCTQVV INSFYIVTRP LAFLNSVINP VFYFLLGDHF RDMLMNQLRH NFKSL TSFS RWAHELLLSF REKAAAVFTL EDFVGDWEQT AAYNLDQVLE QGGVSSLLQN LAVSVTPIQR IVRSGENALK IDIHVI IPY EGLSADQMAQ IEEVFKVVYP VDDHHFKVIL PYGTLVIDGV TPNMLNYFGR PYEGIAVFDG KKITVTGTLW NGNKIID ER LITPDGSMLF RVTINS UniProtKB: Soluble cytochrome b562, Succinate receptor 1 |
-Macromolecule #5: SUCCINIC ACID
| Macromolecule | Name: SUCCINIC ACID / type: ligand / ID: 5 / Number of copies: 1 / Formula: SIN |
|---|---|
| Molecular weight | Theoretical: 118.088 Da |
| Chemical component information |  ChemComp-SIN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 65.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)