[English] 日本語
 Yorodumi
Yorodumi- EMDB-36423: Structure of XBB spike protein (S) dimer-trimer in complex with b... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of XBB spike protein (S) dimer-trimer in complex with bispecific antibody G7-Fc at 3.75 Angstroms resolution. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Antibody / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationMaturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane ...Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / entry receptor-mediated virion attachment to host cell / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / membrane fusion / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / receptor ligand activity / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.75 Å | |||||||||
 Authors Authors | Hao A / Mao Q / Chen Z / Huang J / Sun L | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of XBB spike protein (S) dimer-trimer in complex with bispecific antibody G7-Fc at 3.75 Angstroms resolution. Authors: Hao A / Mao Q / Chen Z / Huang J / Sun L | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36423.map.gz emd_36423.map.gz | 337.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36423-v30.xml emd-36423-v30.xml emd-36423.xml emd-36423.xml | 16.4 KB 16.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36423.png emd_36423.png | 53.2 KB | ||
| Filedesc metadata |  emd-36423.cif.gz emd-36423.cif.gz | 6.3 KB | ||
| Others |  emd_36423_half_map_1.map.gz emd_36423_half_map_1.map.gz emd_36423_half_map_2.map.gz emd_36423_half_map_2.map.gz | 338.1 MB 338.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36423 http://ftp.pdbj.org/pub/emdb/structures/EMD-36423 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36423 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36423 | HTTPS FTP |
-Validation report
| Summary document |  emd_36423_validation.pdf.gz emd_36423_validation.pdf.gz | 689.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_36423_full_validation.pdf.gz emd_36423_full_validation.pdf.gz | 688.9 KB | Display | |
| Data in XML |  emd_36423_validation.xml.gz emd_36423_validation.xml.gz | 17.9 KB | Display | |
| Data in CIF |  emd_36423_validation.cif.gz emd_36423_validation.cif.gz | 21.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36423 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36423 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36423 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36423 | HTTPS FTP |
-Related structure data
| Related structure data |  8jmmMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36423.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36423.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.19 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36423_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
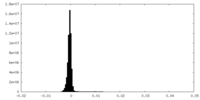
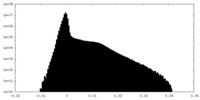
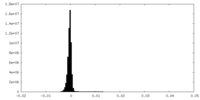
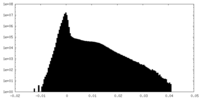
| Density Histograms |
-Half map: #1
| File | emd_36423_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : XBB spike protein (S) in complex with bispecific antibody G7-Fc
| Entire | Name: XBB spike protein (S) in complex with bispecific antibody G7-Fc |
|---|---|
| Components |
|
-Supramolecule #1: XBB spike protein (S) in complex with bispecific antibody G7-Fc
| Supramolecule | Name: XBB spike protein (S) in complex with bispecific antibody G7-Fc type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Details: XBB / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 143.147531 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPMGSLQPLA TLYLLGMLVA SVLAQCVNLI TRTQSYTNSF TRGVYYPDKV FRSSVLHSTQ DLFLPFFSNV TWFHAIHVSG TNGTKRFDN PVLPFNDGVY FASTEKSNII RGWIFGTTLD SKTQSLLIVN NATNVVIKVC EFQFCNDPFL DVYQKNNKSW M ESEFRVYS ...String: MPMGSLQPLA TLYLLGMLVA SVLAQCVNLI TRTQSYTNSF TRGVYYPDKV FRSSVLHSTQ DLFLPFFSNV TWFHAIHVSG TNGTKRFDN PVLPFNDGVY FASTEKSNII RGWIFGTTLD SKTQSLLIVN NATNVVIKVC EFQFCNDPFL DVYQKNNKSW M ESEFRVYS SANNCTFEYV SQPFLMDLEG KEGNFKNLRE FVFKNIDGYF KIYSKHTPIN LERDLPQGFS ALEPLVDLPI GI NITRFQT LLALHRSYLT PGDSSSGWTA GAAAYYVGYL QPRTFLLKYN ENGTITDAVD CALDPLSETK CTLKSFTVEK GIY QTSNFR VQPTESIVRF PNITNLCPFH EVFNATTFAS VYAWNRKRIS NCVADYSVIY NFAPFFAFKC YGVSPTKLND LCFT NVYAD SFVIRGNEVS QIAPGQTGNI ADYNYKLPDD FTGCVIAWNS NKLDSKPSGN YNYLYRLFRK SKLKPFERDI STEIY QAGN KPCNGVAGSN CYSPLQSYGF RPTYGVGHQP YRVVVLSFEL LHAPATVCGP KKSTNLVKNK CVNFNFNGLT GTGVLT ESN KKFLPFQQFG RDIADTTDAV RDPQTLEILD ITPCSFGGVS VITPGTNTSN QVAVLYQGVN CTEVPVAIHA DQLTPTW RV YSTGSNVFQT RAGCLIGAEY VNNSYECDIP IGAGICASYQ TQTKSHGSAS SVASQSIIAY TMSLGAENSV AYSNNSIA I PTNFTISVTT EILPVSMTKT SVDCTMYICG DSTECSNLLL QYGSFCTQLK RALTGIAVEQ DKNTQEVFAQ VKQIYKTPP IKYFGGFNFS QILPDPSKPS KRSPIEDLLF NKVTLADAGF IKQYGDCLGD IAARDLICAQ KFNGLTVLPP LLTDEMIAQY TSALLAGTI TSGWTFGAGP ALQIPFPMQM AYRFNGIGVT QNVLYENQKL IANQFNSAIG KIQDSLSSTP SALGKLQDVV N HNAQALNT LVKQLSSKFG AISSVLNDIL SRLDPPEAEV QIDRLITGRL QSLQTYVTQQ LIRAAEIRAS ANLAATKMSE CV LGQSKRV DFCGKGYHLM SFPQSAPHGV VFLHVTYVPA QEKNFTTAPA ICHDGKAHFP REGVFVSNGT HWFVTQRNFY EPQ IITTDN TFVSGNCDVV IGIVNNTVYD PLQPELDSFK EELDKYFKNH TSPDVDLGDI SGINASVVNI QKEIDRLNEV AKNL NESLI DLQELGKYEQ GSGYIPEAPR DGQAYVRKDG EWVFLSTFLS GLEVLFQGPG GWSHPQFEKG GGSGGGSGGS AWSHP QFEK GGSHHHHHHH H UniProtKB: Spike glycoprotein |
-Macromolecule #2: 7F3 Fv
| Macromolecule | Name: 7F3 Fv / type: protein_or_peptide / ID: 2 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.871873 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EIVLTQSPAT LSLSPGERAT LSCRASQSVS SYLAWYQQKP GQAPRLLIYD ASNRATGIPA RFSGSGSGTD FTLTISSLEP EDFAVYYCQ QRSNWPQYTF GQGTKLEIKG GGGSGGGGSG GGGSEVQLVE SGGGLVKPGR SLRLSCTASG FTFGDYAMSW F RQAPGKGL ...String: EIVLTQSPAT LSLSPGERAT LSCRASQSVS SYLAWYQQKP GQAPRLLIYD ASNRATGIPA RFSGSGSGTD FTLTISSLEP EDFAVYYCQ QRSNWPQYTF GQGTKLEIKG GGGSGGGGSG GGGSEVQLVE SGGGLVKPGR SLRLSCTASG FTFGDYAMSW F RQAPGKGL EWVGFIRSKA YGGTTEYAAS VKGRFTISRD DSKSIAYLQM NSLKTEDTAV YYCTRDRYAR YDILTGLSPA GA DYFYYAM DVWGQGTTVT VSS |
-Macromolecule #3: GW01 Fv
| Macromolecule | Name: GW01 Fv / type: protein_or_peptide / ID: 3 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.330619 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSVLTQPPSA SGTPGQRVTI SCSGSSSNIG SNTVNWYQQL PGTAPKLLIY SNNQRPSGVP DRFSGSKSGT SASLAISGLQ SEDEADYYC AAWDDSLNWV FGGGTKLTVL GGGGSGGGGS GGGGSEVQLV ESGGGVVQPG GSLRLSCAAS GFRFDDHAMH W VRQAPGKG ...String: QSVLTQPPSA SGTPGQRVTI SCSGSSSNIG SNTVNWYQQL PGTAPKLLIY SNNQRPSGVP DRFSGSKSGT SASLAISGLQ SEDEADYYC AAWDDSLNWV FGGGTKLTVL GGGGSGGGGS GGGGSEVQLV ESGGGVVQPG GSLRLSCAAS GFRFDDHAMH W VRQAPGKG LEWVSVISGD GGSTYYADSV KGRFSISRDD SKNSLYLQMN SLRTEDTALY YCAKDRSYGP PDVFNYEYGM DV WGQGTTV TVSSGS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.75 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 539055 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X