+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of hemagglutinin with MSBP | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | glycoproteins / VIRAL PROTEIN | |||||||||
| Biological species |   Influenza A virus Influenza A virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.19 Å | |||||||||
 Authors Authors | Xu Y / Qin Y / Wang L / Zhang Y / Wang Y / Dang S | |||||||||
| Funding support |  Hong Kong, 1 items Hong Kong, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2024 Journal: Commun Biol / Year: 2024Title: Metallo-supramolecular branched polymer protects particles from air-water interface in single-particle cryo-electron microscopy. Authors: Yixin Xu / Yuqi Qin / Lang Wang / Yingyi Zhang / Yufeng Wang / Shangyu Dang /   Abstract: Recent technological breakthroughs in single-particle cryo-electron microscopy (cryo-EM) enable rapid atomic structure determination of biological macromolecules. A major bottleneck in the current ...Recent technological breakthroughs in single-particle cryo-electron microscopy (cryo-EM) enable rapid atomic structure determination of biological macromolecules. A major bottleneck in the current single particle cryo-EM pipeline is the preparation of good quality frozen cryo-EM grids, which is mostly a trial-and-error process. Among many issues, preferred particle orientation and sample damage by air-water interface (AWI) are common practical problems. Here we report a method of applying metallo-supramolecular branched polymer (MSBP) in the cryo-sample preparation for high-resolution single-particle cryo-EM. Our data shows that MSBP keeps a majority of particles away from air-water interface and mitigates preferred orientation as verified by the analyses of apoferritin, hemagglutinin) trimer and various sample proteins. The use of MSBP is a simple method to improve particle distribution for high-resolution structure determination in single-particle cryo-EM. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36314.map.gz emd_36314.map.gz | 92.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36314-v30.xml emd-36314-v30.xml emd-36314.xml emd-36314.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36314.png emd_36314.png | 79.4 KB | ||
| Masks |  emd_36314_msk_1.map emd_36314_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-36314.cif.gz emd-36314.cif.gz | 4 KB | ||
| Others |  emd_36314_half_map_1.map.gz emd_36314_half_map_1.map.gz emd_36314_half_map_2.map.gz emd_36314_half_map_2.map.gz | 95.6 MB 95.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36314 http://ftp.pdbj.org/pub/emdb/structures/EMD-36314 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36314 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36314 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36314.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36314.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
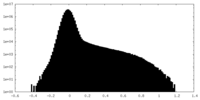
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_36314_msk_1.map emd_36314_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
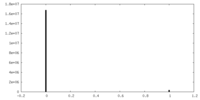
| Density Histograms |
-Half map: #1
| File | emd_36314_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
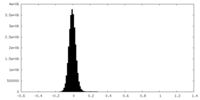
| Density Histograms |
-Half map: #2
| File | emd_36314_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
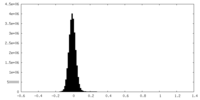
| Density Histograms |
- Sample components
Sample components
-Entire : Hemagglutinin with MSBP
| Entire | Name: Hemagglutinin with MSBP |
|---|---|
| Components |
|
-Supramolecule #1: Hemagglutinin with MSBP
| Supramolecule | Name: Hemagglutinin with MSBP / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Influenza A virus Influenza A virus |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 51.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.3 µm / Nominal magnification: 81000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 3.19 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 231931 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)