[English] 日本語
 Yorodumi
Yorodumi- EMDB-35985: Cryo-EM structure of starch degradation complex of BAM1-LSF1-MDH -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of starch degradation complex of BAM1-LSF1-MDH | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | amylase / starch degradation / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationchloroplast starch grain / starch grain / chloroplast protein-transporting ATPase activity / Ycf2/FtsHi complex / plastid stroma / carbohydrate phosphatase activity / protein import into chloroplast stroma / stromule / chloroplast inner membrane / chloroplast organization ...chloroplast starch grain / starch grain / chloroplast protein-transporting ATPase activity / Ycf2/FtsHi complex / plastid stroma / carbohydrate phosphatase activity / protein import into chloroplast stroma / stromule / chloroplast inner membrane / chloroplast organization / beta-amylase / beta-amylase activity / starch catabolic process / malate dehydrogenase / L-malate dehydrogenase (NAD+) activity / embryo development ending in seed dormancy / malate metabolic process / apoplast / Hydrolases; Acting on ester bonds; Phosphoric-monoester hydrolases / response to water deprivation / chloroplast envelope / plant-type vacuole / plastid / chloroplast stroma / phosphoprotein phosphatase activity / tricarboxylic acid cycle / response to cold / chloroplast / defense response to bacterium / mitochondrion / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Guan ZY / Liu J / Yan JJ | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Plant Cell / Year: 2023 Journal: Plant Cell / Year: 2023Title: The LIKE SEX FOUR 1-malate dehydrogenase complex functions as a scaffold to recruit β-amylase to promote starch degradation. Authors: Jian Liu / Xuecui Wang / Zeyuan Guan / Menglong Wu / Xinyue Wang / Rong Fan / Fei Zhang / Junjun Yan / Yanjun Liu / Delin Zhang / Ping Yin / Junjie Yan /  Abstract: In plant leaves, starch is composed of glucan polymers that accumulate in chloroplasts as the products of photosynthesis during the day; starch is mobilized at night to continuously provide sugars to ...In plant leaves, starch is composed of glucan polymers that accumulate in chloroplasts as the products of photosynthesis during the day; starch is mobilized at night to continuously provide sugars to sustain plant growth and development. Efficient starch degradation requires the involvement of several enzymes, including β-amylase and glucan phosphatase. However, how these enzymes cooperate remains largely unclear. Here, we show that the glucan phosphatase LIKE SEX FOUR 1 (LSF1) interacts with plastid NAD-dependent malate dehydrogenase (MDH) to recruit β-amylase (BAM1), thus reconstituting the BAM1-LSF1-MDH complex. The starch hydrolysis activity of BAM1 drastically increased in the presence of LSF1-MDH in vitro. We determined the structure of the BAM1-LSF1-MDH complex by a combination of cryo-electron microscopy, crosslinking mass spectrometry, and molecular docking. The starch-binding domain of the dual-specificity phosphatase and carbohydrate-binding module of LSF1 was docked in proximity to BAM1, thus facilitating BAM1 access to and hydrolysis of the polyglucans of starch, thus revealing the molecular mechanism by which the LSF1-MDH complex improves the starch degradation activity of BAM1. Moreover, LSF1 is phosphatase inactive, and the enzymatic activity of MDH was dispensable for starch degradation, suggesting nonenzymatic scaffold functions for LSF1-MDH in starch degradation. These findings provide important insights into the precise regulation of starch degradation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35985.map.gz emd_35985.map.gz | 49.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35985-v30.xml emd-35985-v30.xml emd-35985.xml emd-35985.xml | 19.7 KB 19.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35985.png emd_35985.png | 77.3 KB | ||
| Filedesc metadata |  emd-35985.cif.gz emd-35985.cif.gz | 6.7 KB | ||
| Others |  emd_35985_half_map_1.map.gz emd_35985_half_map_1.map.gz emd_35985_half_map_2.map.gz emd_35985_half_map_2.map.gz | 49 MB 49 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35985 http://ftp.pdbj.org/pub/emdb/structures/EMD-35985 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35985 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35985 | HTTPS FTP |
-Validation report
| Summary document |  emd_35985_validation.pdf.gz emd_35985_validation.pdf.gz | 807.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_35985_full_validation.pdf.gz emd_35985_full_validation.pdf.gz | 806.8 KB | Display | |
| Data in XML |  emd_35985_validation.xml.gz emd_35985_validation.xml.gz | 11.9 KB | Display | |
| Data in CIF |  emd_35985_validation.cif.gz emd_35985_validation.cif.gz | 13.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35985 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35985 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35985 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35985 | HTTPS FTP |
-Related structure data
| Related structure data |  8j5dMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35985.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35985.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
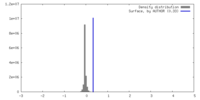
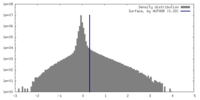
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35985_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
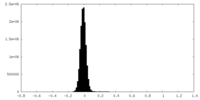
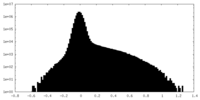
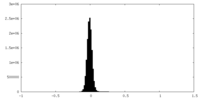
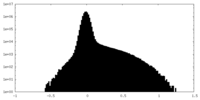
| Density Histograms |
-Half map: #1
| File | emd_35985_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : BAM1-LSF1-MDH complex
| Entire | Name: BAM1-LSF1-MDH complex |
|---|---|
| Components |
|
-Supramolecule #1: BAM1-LSF1-MDH complex
| Supramolecule | Name: BAM1-LSF1-MDH complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Beta-amylase 1, chloroplastic
| Macromolecule | Name: Beta-amylase 1, chloroplastic / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: beta-amylase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 56.111992 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SSGLVPRGSH SDEVDAHMGV PVFVMMPLDS VTMGNTVNRR KAMKASLQAL KSAGVEGIMI DVWWGLVEKE SPGTYNWGG YNELLELAKK LGLKVQAVMS FHQCGGNVGD SVTIPLPQWV VEEVDKDPDL AYTDQWGRRN HEYISLGADT L PVLKGRTP ...String: MGSSHHHHHH SSGLVPRGSH SDEVDAHMGV PVFVMMPLDS VTMGNTVNRR KAMKASLQAL KSAGVEGIMI DVWWGLVEKE SPGTYNWGG YNELLELAKK LGLKVQAVMS FHQCGGNVGD SVTIPLPQWV VEEVDKDPDL AYTDQWGRRN HEYISLGADT L PVLKGRTP VQCYADFMRA FRDNFKHLLG ETIVEIQVGM GPAGELRYPS YPEQEGTWKF PGIGAFQCYD KYSLSSLKAA AE TYGKPEW GSTGPTDAGH YNNWPEDTQF FKKEGGGWNS EYGDFFLSWY SQMLLDHGER ILSSAKSIFE NMGVKISVKI AGI HWHYGT RSHAPELTAG YYNTRFRDGY LPIAQMLARH NAIFNFTCIE MRDHEQPQDA LCAPEKLVNQ VALATLAAEV PLAG ENALP RYDDYAHEQI LKASALNLDQ NNEGEPREMC AFTYLRMNPE LFQADNWGKF VAFVKKMGEG RDSHRCREEV EREAE HFVH VTQPLVQEAA VALTH UniProtKB: Beta-amylase 1, chloroplastic |
-Macromolecule #2: Malate dehydrogenase, chloroplastic
| Macromolecule | Name: Malate dehydrogenase, chloroplastic / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: malate dehydrogenase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 34.120137 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ASYKVAVLGA AGGIGQPLSL LIKMSPLVST LHLYDIANVK GVAADLSHCN TPSQVRDFTG PSELADCLKD VNVVVIPAGV PRKPGMTRD DLFNINANIV KTLVEAVAEN CPNAFIHIIS NPVNSTVPIA AEVLKKKGVY DPKKLFGVTT LDVVRANTFV S QKKNLKLI ...String: ASYKVAVLGA AGGIGQPLSL LIKMSPLVST LHLYDIANVK GVAADLSHCN TPSQVRDFTG PSELADCLKD VNVVVIPAGV PRKPGMTRD DLFNINANIV KTLVEAVAEN CPNAFIHIIS NPVNSTVPIA AEVLKKKGVY DPKKLFGVTT LDVVRANTFV S QKKNLKLI DVDVPVIGGH AGITILPLLS KTKPSVNFTD EEIQELTVRI QNAGTEVVDA KAGAGSATLS MAYAAARFVE SS LRALDGD GDVYECSFVE STLTDLPFFA SRVKIGKNGL EAVIESDLQG LTEYEQKALE ALKVELKASI DKGVAFANKP AAA AAN UniProtKB: Malate dehydrogenase, chloroplastic |
-Macromolecule #3: Phosphoglucan phosphatase LSF1, chloroplastic
| Macromolecule | Name: Phosphoglucan phosphatase LSF1, chloroplastic / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on ester bonds; Phosphoric-monoester hydrolases |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 59.231191 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KMNLNEYMVT LEKPLGIRFA LSADGKIFVH AIKKGSNAEK ARIIMVGDTL KKASDSSGGT LVEIKDFGDT KKMLVEKTGS FSLVLERPF SPFPIQYLLH LSDLDLLYNR GRVSFVTWNK NLLSSNLRAS SQGSGNSGYA AFSSKFFTPQ GWKLLNRQSN S FQSGTKKN ...String: KMNLNEYMVT LEKPLGIRFA LSADGKIFVH AIKKGSNAEK ARIIMVGDTL KKASDSSGGT LVEIKDFGDT KKMLVEKTGS FSLVLERPF SPFPIQYLLH LSDLDLLYNR GRVSFVTWNK NLLSSNLRAS SQGSGNSGYA AFSSKFFTPQ GWKLLNRQSN S FQSGTKKN ILSPPISPLV SVFSEDVPGD GEWGYGNFPL EEYIKALDRS KGELSYNHAL GMRYSKITEQ IYVGSCIQTE ED VENLSEA GITAILNFQG GTEAQNWGID SQSINDACQK SEVLMINYPI KDADSFDLRK KLPLCVGLLL RLLKKNHRVF VTC TTGFDR SSACVIAYLH WMTDTSLHAA YSFVTGLHAC KPDRPAIAWA TWDLIAMVDD GKHDGTPTHS VTFVWNGHEG EEVL LVGDF TGNWKEPIKA THKGGPRFET EVRLTQGKYY YKYIINGDWR HSATSPTERD DRGNTNNIIV VGDVANVRPT IQQPR KDAN IIKVIERVLT ESERFRLAKA ARCIAFSVCP IRLCPKSLEH HHHHH UniProtKB: Phosphoglucan phosphatase LSF1, chloroplastic |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)