+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of dodecameric RdrB cage | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cryoelectron microscopy / adenosine deaminase / IMMUNE SYSTEM | |||||||||
| Function / homology | deaminase activity / Adenosine/adenine deaminase / Metal-dependent hydrolase / Adenosine deaminase Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.58 Å | |||||||||
 Authors Authors | Gao Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Molecular basis of RADAR anti-phage supramolecular assemblies. Authors: Yina Gao / Xiu Luo / Peipei Li / Zhaolong Li / Feng Ye / Songqing Liu / Pu Gao /  Abstract: Adenosine-to-inosine RNA editing has been proposed to be involved in a bacterial anti-phage defense system called RADAR. RADAR contains an adenosine triphosphatase (RdrA) and an adenosine deaminase ...Adenosine-to-inosine RNA editing has been proposed to be involved in a bacterial anti-phage defense system called RADAR. RADAR contains an adenosine triphosphatase (RdrA) and an adenosine deaminase (RdrB). Here, we report cryo-EM structures of RdrA, RdrB, and currently identified RdrA-RdrB complexes in the presence or absence of RNA and ATP. RdrB assembles into a dodecameric cage with catalytic pockets facing outward, while RdrA adopts both autoinhibited tetradecameric and activation-competent heptameric rings. Structural and functional data suggest a model in which RNA is loaded through the bottom section of the RdrA ring and translocated along its inner channel, a process likely coupled with ATP-binding status. Intriguingly, up to twelve RdrA rings can dock one RdrB cage with precise alignments between deaminase catalytic pockets and RNA-translocation channels, indicative of enzymatic coupling of RNA translocation and deamination. Our data uncover an interesting mechanism of enzymatic coupling and anti-phage defense through supramolecular assemblies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34967.map.gz emd_34967.map.gz | 227.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34967-v30.xml emd-34967-v30.xml emd-34967.xml emd-34967.xml | 13.4 KB 13.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34967.png emd_34967.png | 96.8 KB | ||
| Others |  emd_34967_half_map_1.map.gz emd_34967_half_map_1.map.gz emd_34967_half_map_2.map.gz emd_34967_half_map_2.map.gz | 191.4 MB 191.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34967 http://ftp.pdbj.org/pub/emdb/structures/EMD-34967 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34967 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34967 | HTTPS FTP |
-Related structure data
| Related structure data |  8hrcMC  8hr7C  8hr8C  8hr9C  8hraC  8hrbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34967.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34967.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_34967_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
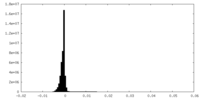
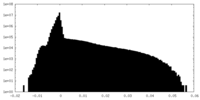
| Density Histograms |
-Half map: #2
| File | emd_34967_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
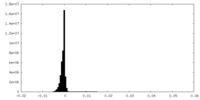
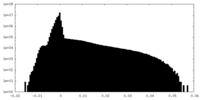
| Density Histograms |
- Sample components
Sample components
-Entire : RdrB
| Entire | Name: RdrB |
|---|---|
| Components |
|
-Supramolecule #1: RdrB
| Supramolecule | Name: RdrB / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Adenosine deaminase
| Macromolecule | Name: Adenosine deaminase / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 92.216953 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MERFLLNSTV LLYRLSTVSL DEVSLDERVE SSVFLAQYEQ ARSLPDHVAK SAWSYLVQQI KQRNMKLGPV AILRLIAEKF IKNEKGGPK IDLPMFSEWQ TLMSRVSCLP IIACHQVFNP GPASQEYSFR WPLYPYHPTV EDYITRECLH ETHQHLNGST S AEECWLDA ...String: MERFLLNSTV LLYRLSTVSL DEVSLDERVE SSVFLAQYEQ ARSLPDHVAK SAWSYLVQQI KQRNMKLGPV AILRLIAEKF IKNEKGGPK IDLPMFSEWQ TLMSRVSCLP IIACHQVFNP GPASQEYSFR WPLYPYHPTV EDYITRECLH ETHQHLNGST S AEECWLDA LKHPEACLRD FEKGWASQEM KQLCAQIDPS LTPRIFKDRL QIACNIREIL CRVAQGVELP EWIASMQNPQ QL ANSTILH NGREYGFATV WPIDDKYSQE SEFCWLTGLL EKWRFNAPEG LERLLWIYLL IQNQYLTLLV QRDDFFGFEQ FQN YTMTEL REETEKSYLS RFKHAHGAGV YSQVRYLEGR FAPKSDPNKM QKLLFSVLRG YWEYLSAHMS MEWVHEKPLT ISQV LDNLE LVEPHGKCVE LALVPHFIKR KPKNGEAYPH ALLFKDLKNQ AAILMDMLKS EPRLTGWIRG VDAAANEMHA PPELF CPLF RVLAKSGIAH FTYHVGEDFP HLISGIRSID DALRFLPLRN GDRLGHCTAI GITPSIWKRS LPLSLSMTKE TRLLDL VFI WRELRSHPEL LRYASDAAIE AVRLAHKVFS LEEEVSITTL DQVFEMRGLL AESEGLLSEL NEPLKPKSLW LEEYERA RE LVKTTGMKRP LKLYKQWLTS DNVRKQRAEY VEVALEYLPD EAVVALQQAV MAKMADRNIA IECPPTSNTR ISQYRNVS E HHIFRWMGLP GEAIEGDVPM SICLGSDDPG IFAADLKSEF YHLFVVLTRK FGLSPADALR KVAEVNENGR IYRFHDVS UniProtKB: Adenosine deaminase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.58 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 373491 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)