[English] 日本語
 Yorodumi
Yorodumi- EMDB-34878: Cryo-EM structure of Mycobacterial Type VII Secretion System Viru... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Mycobacterial Type VII Secretion System Virulence Factor EspB (residues 1-332) | ||||||||||||
 Map data Map data | Cryo-EM structure of Mycobacterial Type VII Secretion System Virulence Factor EspB (residues 1-332) | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Mycobacterium tuberculosis / T7SS / membrane / lipid / LIPID BINDING PROTEIN | ||||||||||||
| Biological species |  | ||||||||||||
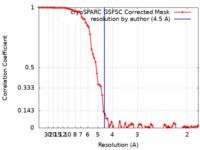
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | ||||||||||||
 Authors Authors | Sengupta N / Padmanaban S / Dutta S | ||||||||||||
| Funding support |  India, 3 items India, 3 items
| ||||||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2023 Journal: J Biol Chem / Year: 2023Title: Cryo-EM reveals the membrane-binding phenomenon of EspB, a virulence factor of the mycobacterial type VII secretion system. Authors: Nayanika Sengupta / Surekha Padmanaban / Somnath Dutta /  Abstract: Mycobacterium tuberculosis (Mtb) utilizes sophisticated machinery called the type VII secretion system to translocate virulence factors across its complex lipid membrane. EspB, a ∼36 kDa secreted ...Mycobacterium tuberculosis (Mtb) utilizes sophisticated machinery called the type VII secretion system to translocate virulence factors across its complex lipid membrane. EspB, a ∼36 kDa secreted substrate of the ESX-1 apparatus, was shown to cause ESAT-6-independent host cell death. Despite the current wealth of high-resolution structural information of the ordered N-terminal domain, the mechanism of EspB-mediated virulence remains poorly characterized. Here, we document EspB interaction with phosphatidic acid (PA) and phosphatidylserine (PS) in the context of membranes, through a biophysical approach including transmission electron microscopy and cryo-EM. We were also able to show PA, PS-dependent conversion of monomers to oligomers at physiological pH. Our data suggest that EspB adheres to biological membranes with limited PA and PS. EM of yeast mitochondria with EspB indicates a mitochondrial membrane-binding property of this ESX-1 substrate. Further, we determined the 3D structures of EspB with and without PA and observed plausible stabilization of the low complexity C-terminal domain in the presence of PA. Collectively, our cryo-EM-based structural and functional studies of EspB provide further insight into the host-Mtb interaction. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34878.map.gz emd_34878.map.gz | 48.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34878-v30.xml emd-34878-v30.xml emd-34878.xml emd-34878.xml | 13 KB 13 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34878_fsc.xml emd_34878_fsc.xml | 9 KB | Display |  FSC data file FSC data file |
| Images |  emd_34878.png emd_34878.png | 195.4 KB | ||
| Filedesc metadata |  emd-34878.cif.gz emd-34878.cif.gz | 3.8 KB | ||
| Others |  emd_34878_half_map_1.map.gz emd_34878_half_map_1.map.gz emd_34878_half_map_2.map.gz emd_34878_half_map_2.map.gz | 40.7 MB 40.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34878 http://ftp.pdbj.org/pub/emdb/structures/EMD-34878 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34878 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34878 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34878.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34878.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of Mycobacterial Type VII Secretion System Virulence Factor EspB (residues 1-332) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.92 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Cryo-EM structure of Mycobacterial Type VII Secretion System...
| File | emd_34878_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of Mycobacterial Type VII Secretion System Virulence Factor EspB (residues 1-332) - half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryo-EM structure of Mycobacterial Type VII Secretion System...
| File | emd_34878_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of Mycobacterial Type VII Secretion System Virulence Factor EspB (residues 1-332) - half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of Mycobacterial Type VII Secretion System Viru...
| Entire | Name: Cryo-EM structure of Mycobacterial Type VII Secretion System Virulence Factor EspB (residues 1-332) |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of Mycobacterial Type VII Secretion System Viru...
| Supramolecule | Name: Cryo-EM structure of Mycobacterial Type VII Secretion System Virulence Factor EspB (residues 1-332) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.75 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)