[English] 日本語
 Yorodumi
Yorodumi- EMDB-34756: F1 domain of FoF1-ATPase from Bacillus PS3, 90 degrees, low ATP -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | F1 domain of FoF1-ATPase from Bacillus PS3, 90 degrees, low ATP | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | ATP synthase F1 ATPase FoF1 / MOTOR PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationproton motive force-driven plasma membrane ATP synthesis / H+-transporting two-sector ATPase / proton-transporting ATP synthase complex / proton-transporting ATP synthase activity, rotational mechanism / ADP binding / ATP binding / plasma membrane Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||||||||
 Authors Authors | Nakano A / Kishikawa J / Mitsuoka K / Yokoyama K | |||||||||||||||
| Funding support |  Japan, 4 items Japan, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Mechanism of ATP hydrolysis dependent rotation of bacterial ATP synthase. Authors: Atsuki Nakano / Jun-Ichi Kishikawa / Kaoru Mitsuoka / Ken Yokoyama /  Abstract: F domain of ATP synthase is a rotary ATPase complex in which rotation of central γ-subunit proceeds in 120° steps against a surrounding αβ fueled by ATP hydrolysis. How the ATP hydrolysis ...F domain of ATP synthase is a rotary ATPase complex in which rotation of central γ-subunit proceeds in 120° steps against a surrounding αβ fueled by ATP hydrolysis. How the ATP hydrolysis reactions occurring in three catalytic αβ dimers are coupled to mechanical rotation is a key outstanding question. Here we describe catalytic intermediates of the F domain in FF synthase from Bacillus PS3 sp. during ATP mediated rotation captured using cryo-EM. The structures reveal that three catalytic events and the first 80° rotation occur simultaneously in F domain when nucleotides are bound at all the three catalytic αβ dimers. The remaining 40° rotation of the complete 120° step is driven by completion of ATP hydrolysis at αβ, and proceeds through three sub-steps (83°, 91°, 101°, and 120°) with three associated conformational intermediates. All sub-steps except for one between 91° and 101° associated with phosphate release, occur independently of the chemical cycle, suggesting that the 40° rotation is largely driven by release of intramolecular strain accumulated by the 80° rotation. Together with our previous results, these findings provide the molecular basis of ATP driven rotation of ATP synthases. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34756.map.gz emd_34756.map.gz | 140.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34756-v30.xml emd-34756-v30.xml emd-34756.xml emd-34756.xml | 22.1 KB 22.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34756_fsc.xml emd_34756_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_34756.png emd_34756.png | 84.3 KB | ||
| Masks |  emd_34756_msk_1.map emd_34756_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-34756.cif.gz emd-34756.cif.gz | 6.6 KB | ||
| Others |  emd_34756_additional_1.map.gz emd_34756_additional_1.map.gz emd_34756_half_map_1.map.gz emd_34756_half_map_1.map.gz emd_34756_half_map_2.map.gz emd_34756_half_map_2.map.gz | 166.6 MB 140.6 MB 140.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34756 http://ftp.pdbj.org/pub/emdb/structures/EMD-34756 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34756 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34756 | HTTPS FTP |
-Validation report
| Summary document |  emd_34756_validation.pdf.gz emd_34756_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_34756_full_validation.pdf.gz emd_34756_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_34756_validation.xml.gz emd_34756_validation.xml.gz | 20 KB | Display | |
| Data in CIF |  emd_34756_validation.cif.gz emd_34756_validation.cif.gz | 26.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34756 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34756 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34756 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34756 | HTTPS FTP |
-Related structure data
| Related structure data |  8hh9MC  8hh1C  8hh2C  8hh3C  8hh4C  8hh5C  8hh6C  8hh7C  8hh8C  8hhaC  8hhbC  8hhcC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34756.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34756.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||
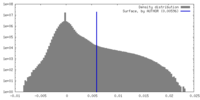
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_34756_msk_1.map emd_34756_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_34756_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34756_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_34756_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : FoF1 from Bacillus PS3
| Entire | Name: FoF1 from Bacillus PS3 |
|---|---|
| Components |
|
-Supramolecule #1: FoF1 from Bacillus PS3
| Supramolecule | Name: FoF1 from Bacillus PS3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 530 KDa |
-Macromolecule #1: ATP synthase subunit alpha
| Macromolecule | Name: ATP synthase subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number: H+-transporting two-sector ATPase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 54.717398 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SIRAEEISAL IKQQIENYES QIQVSDVGTV IQVGDGIARA HGLDNVMSGE LVEFANGVMG MALNLEENNV GIVILGPYTG IKEGDEVRR TGRIMEVPVG EALIGRVVNP LGQPVDGLGP VETTETRPIE SPAPGVMDRR SVHEPLQTGI KAIDALVPIG R GQRELIIG ...String: SIRAEEISAL IKQQIENYES QIQVSDVGTV IQVGDGIARA HGLDNVMSGE LVEFANGVMG MALNLEENNV GIVILGPYTG IKEGDEVRR TGRIMEVPVG EALIGRVVNP LGQPVDGLGP VETTETRPIE SPAPGVMDRR SVHEPLQTGI KAIDALVPIG R GQRELIIG DRQTGKTSVA IDTIINQKDQ NMISIYVAIG QKESTVRTVV ETLRKHGALD YTIVVTASAS QPAPLLFLAP YA GVAMGEY FMYKGKHVLV VYDDLSKQAA AYRELSLLLR RPPGREAYPG DIFYLHSRLL ERAAKLSDAK GGGSLTALPF VET QAGDIS AYIPTNVISI TDGQIFLQSD LFFSGVRPAI NAGLSVSRVG GAAQIKAMKK VAGTLRLDLA AYRELEAFAQ FGSD LDKAT QAKLARGART VEVLKQDLHQ PIPVEKQVLI IYALTRGFLD DIPVEDVRRF EKEFYLFLDQ NGQHLLEHIR TTKDL PNED DLNKAIEAFK KTFVVSQ UniProtKB: ATP synthase subunit alpha |
-Macromolecule #2: ATP synthase subunit beta
| Macromolecule | Name: ATP synthase subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number: H+-transporting two-sector ATPase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 53.424625 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHHHH HMTRGRVIQV MGPVVDVKFE NGHLPAIYNA LKIQHKARNE NEVDIDLTLE VALHLGDDTV RTIAMASTDG LIRGMEVID TGAPISVPVG EVTLGRVFNV LGEPIDLEGD IPADARRDPI HRPAPKFEEL ATEVEILETG IKVVDLLAPY I KGGKIGLF ...String: MHHHHHHHHH HMTRGRVIQV MGPVVDVKFE NGHLPAIYNA LKIQHKARNE NEVDIDLTLE VALHLGDDTV RTIAMASTDG LIRGMEVID TGAPISVPVG EVTLGRVFNV LGEPIDLEGD IPADARRDPI HRPAPKFEEL ATEVEILETG IKVVDLLAPY I KGGKIGLF GGAGVGKTVL IQELIHNIAQ EHGGISVFAG VGERTREGND LYHEMKDSGV ISKTAMVFGQ MNEPPGARMR VA LTGLTMA EYFRDEQGQD VLLFIDNIFR FTQAGSEVSA LLGRMPSAVG YQPTLATEMG QLQERITSTA KGSITSIQAI YVP ADDYTD PAPATTFSHL DATTNLERKL AEMGIYPAVD PLASTSRALA PEIVGEEHYQ VARKVQQTLQ RYKELQDIIA ILGM DELSD EDKLVVHRAR RIQFFLSQNF HVAEQFTGQP GSYVPVKETV RGFKEILEGK YDHLPEDAFR LVGRIEEVVE KAKAM GVEV UniProtKB: ATP synthase subunit beta |
-Macromolecule #3: ATP synthase gamma chain
| Macromolecule | Name: ATP synthase gamma chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 31.728328 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ASLRDIKTRI NATKKTSQIT KAMEMVSTSK LNRAEQNAKS FVPYMEKIQE VVANVALGAG GASHPMLVSR PVKKTGYLVI TSDRGLAGA YNSNVLRLVY QTIQKRHASP DEYAIIVIGR VGLSFFRKRN MPVILDITRL PDQPSFADIK EIARKTVGLF A DGTFDELY ...String: ASLRDIKTRI NATKKTSQIT KAMEMVSTSK LNRAEQNAKS FVPYMEKIQE VVANVALGAG GASHPMLVSR PVKKTGYLVI TSDRGLAGA YNSNVLRLVY QTIQKRHASP DEYAIIVIGR VGLSFFRKRN MPVILDITRL PDQPSFADIK EIARKTVGLF A DGTFDELY MYYNHYVSAI QQEVTERKLL PLTDLAENKQ RTVYEFEPSQ EEILDVLLPQ YAESLIYGAL LDAKASEHAA RM TAMKNAT DNANELIRTL TLSYNRARQA AITQEITEIV AGANALQ UniProtKB: ATP synthase gamma chain |
-Macromolecule #4: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 4 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 5 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: PHOSPHATE ION
| Macromolecule | Name: PHOSPHATE ION / type: ligand / ID: 6 / Number of copies: 2 / Formula: PO4 |
|---|---|
| Molecular weight | Theoretical: 94.971 Da |
| Chemical component information |  ChemComp-PO4: |
-Macromolecule #7: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 7 / Number of copies: 1 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 7653 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.03843 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)