[English] 日本語
 Yorodumi
Yorodumi- EMDB-34406: Local refinement of SARS-CoV-2 Omicron BA.1 Spike glycoprotein in... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Local refinement of SARS-CoV-2 Omicron BA.1 Spike glycoprotein in complex with rabbit monoclonal antibody 1H1 Fab | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | VIRAL PROTEIN-IMMUNE SYSTEM COMPLEX | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.52 Å | |||||||||
 Authors Authors | Guo H / Gao Y / Lu Y / Yang H / Ji X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2023 Journal: Commun Biol / Year: 2023Title: Mechanism of a rabbit monoclonal antibody broadly neutralizing SARS-CoV-2 variants. Authors: Hangtian Guo / Yixuan Yang / Tiantian Zhao / Yuchi Lu / Yan Gao / Tinghan Li / Hang Xiao / Xiaoyu Chu / Le Zheng / Wanting Li / Hao Cheng / Haibin Huang / Yang Liu / Yang Lou / Henry C ...Authors: Hangtian Guo / Yixuan Yang / Tiantian Zhao / Yuchi Lu / Yan Gao / Tinghan Li / Hang Xiao / Xiaoyu Chu / Le Zheng / Wanting Li / Hao Cheng / Haibin Huang / Yang Liu / Yang Lou / Henry C Nguyen / Chao Wu / Yuxin Chen / Haitao Yang / Xiaoyun Ji /  Abstract: Due to the continuous evolution of SARS-CoV-2, the Omicron variant has emerged and exhibits severe immune evasion. The high number of mutations at key antigenic sites on the spike protein has made a ...Due to the continuous evolution of SARS-CoV-2, the Omicron variant has emerged and exhibits severe immune evasion. The high number of mutations at key antigenic sites on the spike protein has made a large number of existing antibodies and vaccines ineffective against this variant. Therefore, it is urgent to develop efficient broad-spectrum neutralizing therapeutic drugs. Here we characterize a rabbit monoclonal antibody (RmAb) 1H1 with broad-spectrum neutralizing potency against Omicron sublineages including BA.1, BA.1.1, BA.2, BA.2.12.1, BA.2.75, BA.3 and BA.4/5. Cryo-electron microscopy (cryo-EM) structure determination of the BA.1 spike-1H1 Fab complexes shows that 1H1 targets a highly conserved region of RBD and avoids most of the circulating Omicron mutations, explaining its broad-spectrum neutralization potency. Our findings indicate 1H1 as a promising RmAb model for designing broad-spectrum neutralizing antibodies and shed light on the development of therapeutic agents as well as effective vaccines against newly emerging variants in the future. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34406.map.gz emd_34406.map.gz | 483.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34406-v30.xml emd-34406-v30.xml emd-34406.xml emd-34406.xml | 20.1 KB 20.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34406.png emd_34406.png | 48.7 KB | ||
| Filedesc metadata |  emd-34406.cif.gz emd-34406.cif.gz | 6.2 KB | ||
| Others |  emd_34406_half_map_1.map.gz emd_34406_half_map_1.map.gz emd_34406_half_map_2.map.gz emd_34406_half_map_2.map.gz | 475.5 MB 475.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34406 http://ftp.pdbj.org/pub/emdb/structures/EMD-34406 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34406 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34406 | HTTPS FTP |
-Related structure data
| Related structure data |  8gzzMC  8h00C  8h01C  8ituC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34406.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34406.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34406_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
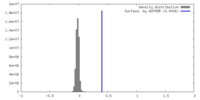
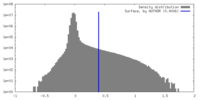
| Density Histograms |
-Half map: #1
| File | emd_34406_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
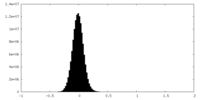
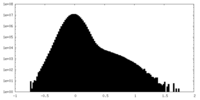
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 Omicron spike glycoprotein complex with 55A8 Fab
| Entire | Name: SARS-CoV-2 Omicron spike glycoprotein complex with 55A8 Fab |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 Omicron spike glycoprotein complex with 55A8 Fab
| Supramolecule | Name: SARS-CoV-2 Omicron spike glycoprotein complex with 55A8 Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: Omicron spike glycoprotein
| Supramolecule | Name: Omicron spike glycoprotein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 Details: SARS-CoV-2 Omicron BA.1 variant spike protein receptor binding domain |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: 1H1 Fab
| Supramolecule | Name: 1H1 Fab / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 Details: Fab fragment generated by proteolytic cleavage of IgG antibody |
|---|
-Macromolecule #1: Spike protein S1
| Macromolecule | Name: Spike protein S1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.065934 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NLCPFDEVFN ATRFASVYAW NRKRISNCVA DYSVLYNLAP FFTFKCYGVS PTKLNDLCFT NVYADSFVIR GDEVRQIAPG QTGNIADYN YKLPDDFTGC VIAWNSNKLD SKVSGNYNYL YRLFRKSNLK PFERDISTEI YQAGNKPCNG VAGFNCYFPL R SYSFRPTY ...String: NLCPFDEVFN ATRFASVYAW NRKRISNCVA DYSVLYNLAP FFTFKCYGVS PTKLNDLCFT NVYADSFVIR GDEVRQIAPG QTGNIADYN YKLPDDFTGC VIAWNSNKLD SKVSGNYNYL YRLFRKSNLK PFERDISTEI YQAGNKPCNG VAGFNCYFPL R SYSFRPTY GVGHQPYRVV VLSFELLHAP ATVCGP UniProtKB: Spike glycoprotein |
-Macromolecule #2: rabbit monoclonal antibody 1H1 Fab light chain
| Macromolecule | Name: rabbit monoclonal antibody 1H1 Fab light chain / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.739082 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIVMTQTPAS VSEPVGGTVT IKCQASESIS NWLAWYQQKP GQPPKLLIYA AFTLASGVPS RFKGSGSGTQ FTLTINGVEC ADAATYYCQ QTYSSRDVDN VFGGGTEVVV KG |
-Macromolecule #3: rabbit monoclonal antibody 1H1 Fab heavy chain
| Macromolecule | Name: rabbit monoclonal antibody 1H1 Fab heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.863201 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSLEESGGDL VKPGASLTLT CTASGFSFSS GYDMCWVRQA PGKGLEWIAC IGTGSSGNIY YASWAKGRFT ISKTSSTTVT LQMTSLTAA DTATYFCARD DADYAGPDYF NLWGPGTLVT VSS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)