[English] 日本語
 Yorodumi
Yorodumi- EMDB-33595: Cryo-electron tomography of fixed SARS-CoV-2 D614 strain (aligned... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-electron tomography of fixed SARS-CoV-2 D614 strain (aligned tilt series). | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / Virus | |||||||||
| Biological species |  | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Song Y / Li S | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2020 Journal: Cell / Year: 2020Title: Molecular Architecture of the SARS-CoV-2 Virus. Authors: Hangping Yao / Yutong Song / Yong Chen / Nanping Wu / Jialu Xu / Chujie Sun / Jiaxing Zhang / Tianhao Weng / Zheyuan Zhang / Zhigang Wu / Linfang Cheng / Danrong Shi / Xiangyun Lu / Jianlin ...Authors: Hangping Yao / Yutong Song / Yong Chen / Nanping Wu / Jialu Xu / Chujie Sun / Jiaxing Zhang / Tianhao Weng / Zheyuan Zhang / Zhigang Wu / Linfang Cheng / Danrong Shi / Xiangyun Lu / Jianlin Lei / Max Crispin / Yigong Shi / Lanjuan Li / Sai Li /   Abstract: SARS-CoV-2 is an enveloped virus responsible for the COVID-19 pandemic. Despite recent advances in the structural elucidation of SARS-CoV-2 proteins, the detailed architecture of the intact virus ...SARS-CoV-2 is an enveloped virus responsible for the COVID-19 pandemic. Despite recent advances in the structural elucidation of SARS-CoV-2 proteins, the detailed architecture of the intact virus remains to be unveiled. Here we report the molecular assembly of the authentic SARS-CoV-2 virus using cryoelectron tomography (cryo-ET) and subtomogram averaging (STA). Native structures of the S proteins in pre- and postfusion conformations were determined to average resolutions of 8.7-11 Å. Compositions of the N-linked glycans from the native spikes were analyzed by mass spectrometry, which revealed overall processing states of the native glycans highly similar to that of the recombinant glycoprotein glycans. The native conformation of the ribonucleoproteins (RNPs) and their higher-order assemblies were revealed. Overall, these characterizations revealed the architecture of the SARS-CoV-2 virus in exceptional detail and shed light on how the virus packs its ∼30-kb-long single-segmented RNA in the ∼80-nm-diameter lumen. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33595.map.gz emd_33595.map.gz | 189.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33595-v30.xml emd-33595-v30.xml emd-33595.xml emd-33595.xml | 9.7 KB 9.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33595.png emd_33595.png | 332 KB | ||
| Filedesc metadata |  emd-33595.cif.gz emd-33595.cif.gz | 4.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33595 http://ftp.pdbj.org/pub/emdb/structures/EMD-33595 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33595 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33595 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33595.map.gz / Format: CCP4 / Size: 230.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33595.map.gz / Format: CCP4 / Size: 230.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.44 Å | ||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Severe acute respiratory syndrome coronavirus 2
| Entire | Name:  |
|---|---|
| Components |
|
-Supramolecule #1: Severe acute respiratory syndrome coronavirus 2
| Supramolecule | Name: Severe acute respiratory syndrome coronavirus 2 / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 2697049 Sci species name: Severe acute respiratory syndrome coronavirus 2 Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE / Instrument: GATAN CRYOPLUNGE 3 |
| Sectioning | Other: NO SECTIONING |
| Fiducial marker | Manufacturer: Aurion / Diameter: 10 nm |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Average exposure time: 0.213 sec. / Average electron dose: 3.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 64000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Number images used: 328 |
|---|
 Movie
Movie Controller
Controller


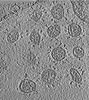






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
















