+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of MrgD-Gi complex with beta-alanine | |||||||||
 Map data Map data | post-process map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / Complex / SIGNALING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationActivation of the phototransduction cascade / Olfactory Signaling Pathway / angiotensin-mediated vasodilation involved in regulation of systemic arterial blood pressure / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Prostacyclin signalling through prostacyclin receptor / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G alpha (z) signalling events / Glucagon-type ligand receptors / G beta:gamma signalling through PI3Kgamma ...Activation of the phototransduction cascade / Olfactory Signaling Pathway / angiotensin-mediated vasodilation involved in regulation of systemic arterial blood pressure / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Prostacyclin signalling through prostacyclin receptor / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G alpha (z) signalling events / Glucagon-type ligand receptors / G beta:gamma signalling through PI3Kgamma / G beta:gamma signalling through CDC42 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Adrenaline,noradrenaline inhibits insulin secretion / ADP signalling through P2Y purinoceptor 12 / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / G beta:gamma signalling through BTK / Thromboxane signalling through TP receptor / Thrombin signalling through proteinase activated receptors (PARs) / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / G alpha (s) signalling events / Ca2+ pathway / G alpha (12/13) signalling events / Extra-nuclear estrogen signaling / G alpha (q) signalling events / Vasopressin regulates renal water homeostasis via Aquaporins / GPER1 signaling / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / ADP signalling through P2Y purinoceptor 1 / G alpha (i) signalling events / G protein-coupled peptide receptor activity / spectrin binding / alkylglycerophosphoethanolamine phosphodiesterase activity / phototransduction, visible light / photoreceptor outer segment / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration / Adenylate cyclase inhibitory pathway / D2 dopamine receptor binding / response to prostaglandin E / cardiac muscle cell apoptotic process / adenylate cyclase regulator activity / G protein-coupled serotonin receptor binding / adenylate cyclase-inhibiting serotonin receptor signaling pathway / cellular response to forskolin / regulation of mitotic spindle organization / Regulation of insulin secretion / positive regulation of cholesterol biosynthetic process / electron transport chain / negative regulation of insulin secretion / G protein-coupled receptor binding / response to peptide hormone / G protein-coupled receptor activity / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / centriolar satellite / G-protein beta/gamma-subunit complex binding / adenylate cyclase-activating G protein-coupled receptor signaling pathway / ADP signalling through P2Y purinoceptor 12 / Adrenaline,noradrenaline inhibits insulin secretion / GDP binding / G alpha (z) signalling events / ADORA2B mediated anti-inflammatory cytokines production / GPER1 signaling / G-protein beta-subunit binding / heterotrimeric G-protein complex / myelin sheath / sensory perception of taste / retina development in camera-type eye / G protein activity / GTPase binding / fibroblast proliferation / midbody / cell cortex / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / cellular response to hypoxia / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / periplasmic space / electron transfer activity / Extra-nuclear estrogen signaling / cell population proliferation / ciliary basal body / G protein-coupled receptor signaling pathway / iron ion binding / lysosomal membrane / cell division / GTPase activity / heme binding / synapse / centrosome / GTP binding / protein-containing complex binding / nucleolus / magnesium ion binding / Golgi apparatus Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
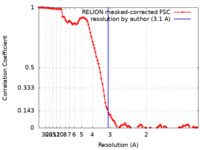
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Suzuki S / Iida M / Kawamoto A / Oshima A | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2022 Journal: Commun Biol / Year: 2022Title: Structural insight into the activation mechanism of MrgD with heterotrimeric Gi-protein revealed by cryo-EM. Authors: Shota Suzuki / Momoko Iida / Yoko Hiroaki / Kotaro Tanaka / Akihiro Kawamoto / Takayuki Kato / Atsunori Oshima /  Abstract: MrgD, a member of the Mas-related G protein-coupled receptor (MRGPR) family, has high basal activity for Gi activation. It recognizes endogenous ligands, such as β-alanine, and is involved in pain ...MrgD, a member of the Mas-related G protein-coupled receptor (MRGPR) family, has high basal activity for Gi activation. It recognizes endogenous ligands, such as β-alanine, and is involved in pain and itch signaling. The lack of a high-resolution structure for MrgD hinders our understanding of whether its activation is ligand-dependent or constitutive. Here, we report two cryo-EM structures of the MrgD-Gi complex in the β-alanine-bound and apo states at 3.1 Å and 2.8 Å resolution, respectively. These structures show that β-alanine is bound to a shallow pocket at the extracellular domains. The extracellular half of the sixth transmembrane helix undergoes a significant movement and is tightly packed into the third transmembrane helix through hydrophobic residues, creating the active form. Our structures demonstrate a structural basis for the characteristic ligand recognition of MrgD. These findings provide a framework to guide drug designs targeting the MrgD receptor. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33554.map.gz emd_33554.map.gz | 9.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33554-v30.xml emd-33554-v30.xml emd-33554.xml emd-33554.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33554_fsc.xml emd_33554_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_33554.png emd_33554.png | 121 KB | ||
| Masks |  emd_33554_msk_1.map emd_33554_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-33554.cif.gz emd-33554.cif.gz | 7.3 KB | ||
| Others |  emd_33554_half_map_1.map.gz emd_33554_half_map_1.map.gz emd_33554_half_map_2.map.gz emd_33554_half_map_2.map.gz | 98.4 MB 98.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33554 http://ftp.pdbj.org/pub/emdb/structures/EMD-33554 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33554 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33554 | HTTPS FTP |
-Validation report
| Summary document |  emd_33554_validation.pdf.gz emd_33554_validation.pdf.gz | 857.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33554_full_validation.pdf.gz emd_33554_full_validation.pdf.gz | 856.9 KB | Display | |
| Data in XML |  emd_33554_validation.xml.gz emd_33554_validation.xml.gz | 18.8 KB | Display | |
| Data in CIF |  emd_33554_validation.cif.gz emd_33554_validation.cif.gz | 24.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33554 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33554 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33554 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33554 | HTTPS FTP |
-Related structure data
| Related structure data |  7y12MC  7y13C  7y14C  7y15C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
| EM raw data |  EMPIAR-11073 (Title: Cryo-EM structure of MrgD-Gi complex with beta-alanine EMPIAR-11073 (Title: Cryo-EM structure of MrgD-Gi complex with beta-alanineData size: 4.1 TB Data #1: K3 movies for MrgD-Gi complex with beta-alanine [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33554.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33554.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | post-process map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.675 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_33554_msk_1.map emd_33554_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_33554_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_33554_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : beta-alanine bound MrgD-Gi complex with the scFv16
+Supramolecule #1: beta-alanine bound MrgD-Gi complex with the scFv16
+Supramolecule #2: G(i) subunit alpha-1,Mas-related G-protein coupled receptor member D
+Supramolecule #3: subunit beta-1,subunit gamma-2
+Supramolecule #4: scFV16
+Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-1
+Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
+Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
+Macromolecule #4: Soluble cytochrome b562,Mas-related G-protein coupled receptor me...
+Macromolecule #5: scFV16
+Macromolecule #6: BETA-ALANINE
+Macromolecule #7: PALMITIC ACID
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot time 3 seconds blot force 5. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: The Microscope implicated Cs corrector. |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 0.7000000000000001 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)