[English] 日本語
 Yorodumi
Yorodumi- EMDB-33415: Cryo-EM structure of SARS-CoV-2 spike protein in complex with nan... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
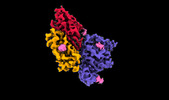
| Title | Cryo-EM structure of SARS-CoV-2 spike protein in complex with nanobody C5G2 | |||||||||
 Map data Map data | Cryo-EM density map of SARS-CoV-2 spike protein in complex with C5G2 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.51 Å | |||||||||
 Authors Authors | Liu L / Sun H / Jiang Y / Liu X / Zhao D / Zheng Q / Li S / Xia N | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: J Nanobiotechnology / Year: 2022 Journal: J Nanobiotechnology / Year: 2022Title: A potent synthetic nanobody with broad-spectrum activity neutralizes SARS-CoV-2 virus and the Omicron variant BA.1 through a unique binding mode. Authors: Dongping Zhao / Liqin Liu / Xinlin Liu / Jinlei Zhang / Yuqing Yin / Linli Luan / Dingwen Jiang / Xiong Yang / Lei Li / Hualong Xiong / Dongming Xing / Qingbing Zheng / Ningshao Xia / Yuyong ...Authors: Dongping Zhao / Liqin Liu / Xinlin Liu / Jinlei Zhang / Yuqing Yin / Linli Luan / Dingwen Jiang / Xiong Yang / Lei Li / Hualong Xiong / Dongming Xing / Qingbing Zheng / Ningshao Xia / Yuyong Tao / Shaowei Li / Haiming Huang /  Abstract: The major challenge to controlling the COVID pandemic is the rapid mutation rate of the SARS-CoV-2 virus, leading to the escape of the protection of vaccines and most of the neutralizing antibodies ...The major challenge to controlling the COVID pandemic is the rapid mutation rate of the SARS-CoV-2 virus, leading to the escape of the protection of vaccines and most of the neutralizing antibodies to date. Thus, it is essential to develop neutralizing antibodies with broad-spectrum activity targeting multiple SARS-CoV-2 variants. Here, we report a synthetic nanobody (named C5G2) obtained by phage display and subsequent antibody engineering. C5G2 has a single-digit nanomolar binding affinity to the RBD domain and inhibits its binding to ACE2 with an IC of 3.7 nM. Pseudovirus assays indicated that monovalent C5G2 could protect the cells from infection with SARS-CoV-2 wild-type virus and most of the viruses of concern, i.e., Alpha, Beta, Gamma and Omicron variants. Strikingly, C5G2 has the highest potency against Omicron BA.1 among all the variants, with an IC of 4.9 ng/mL. The cryo-EM structure of C5G2 in complex with the spike trimer showed that C5G2 binds to RBD mainly through its CDR3 at a conserved region that does not overlap with the ACE2 binding surface. Additionally, C5G2 binds simultaneously to the neighboring NTD domain of the spike trimer through the same CDR3 loop, which may further increase its potency against viral infection. Third, the steric hindrance caused by FR2 of C5G2 could inhibit the binding of ACE2 to RBD as well. Thus, this triple-function nanobody may serve as an effective drug for prophylaxis and therapy against Omicron as well as future variants. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33415.map.gz emd_33415.map.gz | 483.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33415-v30.xml emd-33415-v30.xml emd-33415.xml emd-33415.xml | 12.7 KB 12.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33415.png emd_33415.png | 38.2 KB | ||
| Others |  emd_33415_half_map_1.map.gz emd_33415_half_map_1.map.gz emd_33415_half_map_2.map.gz emd_33415_half_map_2.map.gz | 475.5 MB 475.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33415 http://ftp.pdbj.org/pub/emdb/structures/EMD-33415 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33415 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33415 | HTTPS FTP |
-Validation report
| Summary document |  emd_33415_validation.pdf.gz emd_33415_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33415_full_validation.pdf.gz emd_33415_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_33415_validation.xml.gz emd_33415_validation.xml.gz | 18.5 KB | Display | |
| Data in CIF |  emd_33415_validation.cif.gz emd_33415_validation.cif.gz | 21.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33415 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33415 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33415 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33415 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33415.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33415.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM density map of SARS-CoV-2 spike protein in complex with C5G2 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.778 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Cryo-EM density map of SARS-CoV-2 spike protein in complex with C5G2
| File | emd_33415_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM density map of SARS-CoV-2 spike protein in complex with C5G2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryo-EM density map of SARS-CoV-2 spike protein in complex with C5G2
| File | emd_33415_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM density map of SARS-CoV-2 spike protein in complex with C5G2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of SARS-CoV-2 spike protein in complex with nan...
| Entire | Name: Cryo-EM structure of SARS-CoV-2 spike protein in complex with nanobody C5G2 |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of SARS-CoV-2 spike protein in complex with nan...
| Supramolecule | Name: Cryo-EM structure of SARS-CoV-2 spike protein in complex with nanobody C5G2 type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.51 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 120400 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X