+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a class T GPCR in active state | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | G protein-coupled receptor / taste type 2 receptors / cryo-electron microscopy structure / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationbitter taste receptor activity / detection of chemical stimulus involved in sensory perception of bitter taste / Class C/3 (Metabotropic glutamate/pheromone receptors) / cellulase activity / ciliary membrane / beta-glucosidase activity / cellulose catabolic process / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade ...bitter taste receptor activity / detection of chemical stimulus involved in sensory perception of bitter taste / Class C/3 (Metabotropic glutamate/pheromone receptors) / cellulase activity / ciliary membrane / beta-glucosidase activity / cellulose catabolic process / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G protein-coupled acetylcholine receptor signaling pathway / G-protein activation / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through CDC42 / Glucagon signaling in metabolic regulation / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADP signalling through P2Y purinoceptor 1 / cellular response to catecholamine stimulus / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / adenylate cyclase-activating dopamine receptor signaling pathway / GPER1 signaling / Inactivation, recovery and regulation of the phototransduction cascade / cellular response to prostaglandin E stimulus / G-protein beta-subunit binding / heterotrimeric G-protein complex / G alpha (12/13) signalling events / sensory perception of taste / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / retina development in camera-type eye / GTPase binding / Ca2+ pathway / fibroblast proliferation / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Ras protein signal transduction / Extra-nuclear estrogen signaling / cell population proliferation / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase activity / synapse / protein-containing complex binding / cell surface / signal transduction / extracellular exosome / extracellular region / membrane / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.01 Å | |||||||||||||||
 Authors Authors | Liu ZJ / Hua T / Xu WX / Wu LJ | |||||||||||||||
| Funding support |  China, 4 items China, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Structural basis for strychnine activation of human bitter taste receptor TAS2R46. Authors: Weixiu Xu / Lijie Wu / Shenhui Liu / Xiao Liu / Xiaoling Cao / Cui Zhou / Jinyi Zhang / You Fu / Yu Guo / Yiran Wu / Qiwen Tan / Ling Wang / Junlin Liu / Longquan Jiang / Zhongbo Fan / Yuan ...Authors: Weixiu Xu / Lijie Wu / Shenhui Liu / Xiao Liu / Xiaoling Cao / Cui Zhou / Jinyi Zhang / You Fu / Yu Guo / Yiran Wu / Qiwen Tan / Ling Wang / Junlin Liu / Longquan Jiang / Zhongbo Fan / Yuan Pei / Jingyi Yu / Jianjun Cheng / Suwen Zhao / Xiaojiang Hao / Zhi-Jie Liu / Tian Hua /  Abstract: Taste sensing is a sophisticated chemosensory process, and bitter taste perception is mediated by type 2 taste receptors (TAS2Rs), or class T G protein-coupled receptors. Understanding the detailed ...Taste sensing is a sophisticated chemosensory process, and bitter taste perception is mediated by type 2 taste receptors (TAS2Rs), or class T G protein-coupled receptors. Understanding the detailed molecular mechanisms behind taste sensation is hindered by a lack of experimental receptor structures. Here, we report the cryo-electron microscopy structures of human TAS2R46 complexed with chimeric mini-G protein gustducin, in both strychnine-bound and apo forms. Several features of TAS2R46 are disclosed, including distinct receptor structures that compare with known GPCRs, a new "toggle switch," activation-related motifs, and precoupling with mini-G protein gustducin. Furthermore, the dynamic extracellular and more-static intracellular parts of TAS2R46 suggest possible diverse ligand-recognition and activation processes. This study provides a basis for further exploration of other bitter taste receptors and their therapeutic applications. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33366.map.gz emd_33366.map.gz | 57 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33366-v30.xml emd-33366-v30.xml emd-33366.xml emd-33366.xml | 24.8 KB 24.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33366.png emd_33366.png | 47 KB | ||
| Filedesc metadata |  emd-33366.cif.gz emd-33366.cif.gz | 7.5 KB | ||
| Others |  emd_33366_half_map_1.map.gz emd_33366_half_map_1.map.gz emd_33366_half_map_2.map.gz emd_33366_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33366 http://ftp.pdbj.org/pub/emdb/structures/EMD-33366 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33366 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33366 | HTTPS FTP |
-Validation report
| Summary document |  emd_33366_validation.pdf.gz emd_33366_validation.pdf.gz | 736.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33366_full_validation.pdf.gz emd_33366_full_validation.pdf.gz | 736.4 KB | Display | |
| Data in XML |  emd_33366_validation.xml.gz emd_33366_validation.xml.gz | 12 KB | Display | |
| Data in CIF |  emd_33366_validation.cif.gz emd_33366_validation.cif.gz | 14.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33366 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33366 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33366 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33366 | HTTPS FTP |
-Related structure data
| Related structure data |  7xp6MC  7xp4C  7xp5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33366.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33366.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_33366_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_33366_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of GPCR and G protein
| Entire | Name: Complex of GPCR and G protein |
|---|---|
| Components |
|
-Supramolecule #1: Complex of GPCR and G protein
| Supramolecule | Name: Complex of GPCR and G protein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(t) subunit alpha-3
| Macromolecule | Name: Guanine nucleotide-binding protein G(t) subunit alpha-3 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 30.583334 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDYKDDDDKE NLYFQSNSKT EDQRNEEKAQ REANKKIEKQ LQKDKQVYRA THRLLLLGAD NSGKSTIVKQ MRILHGGSGG SGGTSGIFE TKFQVDKVNF HMFDVGGQRD ERRKWIQCFN DVTAIIFVVD SSDYNRLQEA LNDFKSIWNN RWLRTISVIL F LNKQDLLA ...String: MDYKDDDDKE NLYFQSNSKT EDQRNEEKAQ REANKKIEKQ LQKDKQVYRA THRLLLLGAD NSGKSTIVKQ MRILHGGSGG SGGTSGIFE TKFQVDKVNF HMFDVGGQRD ERRKWIQCFN DVTAIIFVVD SSDYNRLQEA LNDFKSIWNN RWLRTISVIL F LNKQDLLA EKVLAGKSKI EDYFPEFARY TTPEDATPEP GEDPRVTRAK YFIRDEFLRI STASGDGRHY CYPHFTCAVD TQ NVKFVFD AVTDIIIKEN LKDCGLF |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.728426 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL ...String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD INAICFFPNG NA FATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAGHDNRVSC LGV TDDGMA VATGSWDSFL KIWNGSSGGG GSGGGGSSGV SGWRLFKKIS UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Nanobody 35
| Macromolecule | Name: Nanobody 35 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 15.271938 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQVQLQESGG GLVQPGGSLR LSCAASGFTF SNYKMNWVRQ APGKGLEWVS DISQSGASIS YTGSVKGRFT ISRDNAKNTL YLQMNSLKP EDTAVYYCAR CPAPFTRDCF DVTSTTYAYR GQGTQVTVSS HHHHHHEPEA |
-Macromolecule #5: Endoglucanase H,Taste receptor type 2 member 46,Endoglucanase H,T...
| Macromolecule | Name: Endoglucanase H,Taste receptor type 2 member 46,Endoglucanase H,Taste receptor type 2 member 46,Bitter taste receptor T2R46 type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 91.116266 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDAHHHHHH HHHHENLYFQ SGRAMASNYN SGLKIGAWVG TQPSESAIKS FQELQGRKLD IVHQFINWS TDFSWVRPYA DAVYNNGSIL MITWEPWEYN TVDIKNGKAD AYITRMAQDM KAYGKEIWLR PLHAANGDWY P WAIGYSSR ...String: MKTIIALSYI FCLVFADYKD DDDAHHHHHH HHHHENLYFQ SGRAMASNYN SGLKIGAWVG TQPSESAIKS FQELQGRKLD IVHQFINWS TDFSWVRPYA DAVYNNGSIL MITWEPWEYN TVDIKNGKAD AYITRMAQDM KAYGKEIWLR PLHAANGDWY P WAIGYSSR VNTNETYIAA FRHIVDIFRA NGATNVKWVF NVNCDNVGNG TSYLGHYPGD NYVDYTSIDG YNWGTTQSWG SQ WQSFDQV FSRAYQALAS INKPIIIAEF ASAEIGGNKA RWITEAYNSI RTSYNKVIAA VWFHENKETD WRINSSPEAL AAY REAIGA ITFLPIIFSI LIVVTFVIGN FANGFIALVN SIEWFKRQKI SFADQILTAL AVSRVGLLWV LVLNWYATEL NPAF NSIEV RITAYNVWAV INHFSNWLAT SLSIFYLLKI ANFSNLIFLH LKRRVKSVVL VILLGPLLFL VCHLFVINMN QIIWT KEYE GNMTWKIKLR SAMYLSNTTV TILANLVPFT LTLISFLLLI CSLCKHLKKM QLHGKGSQDP SMKVHIKALQ TVTSFL LLC AIYFLSIIMS VWSFESLENK PVFMFCEAIA FSYPSTHPFI LIWGNKKLKQ TFLSVLWHVR YWVKGEKPSS SGSGSSG SG SSVFTLEDFV GDWEQTAAYN LDQVLEQGGV SSLLQNLAVS VTPIQRIVRS GENALKIDIH VIIPYEGLSA DQMAQIEE V FKVVYPVDDH HFKVILPYGT LVIDGVTPNM LNYFGRPYEG IAVFDGKKIT VTGTLWNGNK IIDERLITPD GSMLFRVTI NS UniProtKB: Endoglucanase H, Taste receptor type 2 member 46 |
-Macromolecule #6: STRYCHNINE
| Macromolecule | Name: STRYCHNINE / type: ligand / ID: 6 / Number of copies: 1 / Formula: SY9 |
|---|---|
| Molecular weight | Theoretical: 334.412 Da |
| Chemical component information | 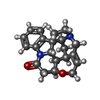 ChemComp-SY9: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK II |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-7xp6: |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)