[English] 日本語
 Yorodumi
Yorodumi- EMDB-33204: S-ECD (Omicron) in complex with PD of ACE2 focused on RBD_PD sub-... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | S-ECD (Omicron) in complex with PD of ACE2 focused on RBD_PD sub-complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Li YN / Shen YP / Zhang YY / Yan RH | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: iScience / Year: 2022 Journal: iScience / Year: 2022Title: Structural and functional analysis of an inter-Spike bivalent neutralizing antibody against SARS-CoV-2 variants. Authors: Yaning Li / Qing Fan / Bing Zhou / Yaping Shen / Yuanyuan Zhang / Lin Cheng / Furong Qi / Shuo Song / Yingying Guo / Renhong Yan / Bin Ju / Zheng Zhang /  Abstract: The different variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) have attracted most public concern because they caused "wave and wave" COVID-19 pandemic. The initial step of ...The different variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) have attracted most public concern because they caused "wave and wave" COVID-19 pandemic. The initial step of viral infection is mediated by the SARS-CoV-2 Spike (S) protein, which mediates the receptor recognition and membrane fusion between virus and host cells. Neutralizing antibodies (nAbs) targeting the S protein of SARS-CoV-2 have become promising candidates for clinical intervention strategy, while multiple studies have shown that different variants have enhanced infectivity and antibody resistance. Here, we explore the structure and function of STS165, a broadly inter-Spike bivalent nAb against SARS-CoV-2 variants and even SARS-CoV, contributing to further understanding of the working mechanism of nAbs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33204.map.gz emd_33204.map.gz | 86 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33204-v30.xml emd-33204-v30.xml emd-33204.xml emd-33204.xml | 13.7 KB 13.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33204.png emd_33204.png | 19.9 KB | ||
| Others |  emd_33204_half_map_1.map.gz emd_33204_half_map_1.map.gz emd_33204_half_map_2.map.gz emd_33204_half_map_2.map.gz | 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33204 http://ftp.pdbj.org/pub/emdb/structures/EMD-33204 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33204 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33204 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33204.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33204.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_33204_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
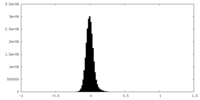
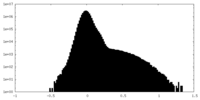
| Density Histograms |
-Half map: #2
| File | emd_33204_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
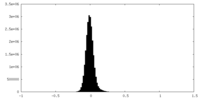
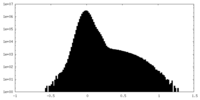
| Density Histograms |
- Sample components
Sample components
-Entire : S-ECD (Omicron) in complex with PD of ACE2
| Entire | Name: S-ECD (Omicron) in complex with PD of ACE2 |
|---|---|
| Components |
|
-Supramolecule #1: S-ECD (Omicron) in complex with PD of ACE2
| Supramolecule | Name: S-ECD (Omicron) in complex with PD of ACE2 / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: S-ECD (Omicron)
| Supramolecule | Name: S-ECD (Omicron) / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: ACE2
| Supramolecule | Name: ACE2 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.0.6) / Number images used: 65532 |
|---|---|
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)