[English] 日本語
 Yorodumi
Yorodumi- EMDB-33195: The Cryo-EM structure of SARS-CoV-2 Omicron 3-RBD up spike protei... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The Cryo-EM structure of SARS-CoV-2 Omicron 3-RBD up spike protein complexed with ZCB11 fab | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.56 Å | |||||||||
 Authors Authors | Hang L / Dang S | |||||||||
| Funding support |  Hong Kong, 1 items Hong Kong, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: A broadly neutralizing antibody protects Syrian hamsters against SARS-CoV-2 Omicron challenge. Authors: Biao Zhou / Runhong Zhou / Bingjie Tang / Jasper Fuk-Woo Chan / Mengxiao Luo / Qiaoli Peng / Shuofeng Yuan / Hang Liu / Bobo Wing-Yee Mok / Bohao Chen / Pui Wang / Vincent Kwok-Man Poon / ...Authors: Biao Zhou / Runhong Zhou / Bingjie Tang / Jasper Fuk-Woo Chan / Mengxiao Luo / Qiaoli Peng / Shuofeng Yuan / Hang Liu / Bobo Wing-Yee Mok / Bohao Chen / Pui Wang / Vincent Kwok-Man Poon / Hin Chu / Chris Chung-Sing Chan / Jessica Oi-Ling Tsang / Chris Chun-Yiu Chan / Ka-Kit Au / Hiu-On Man / Lu Lu / Kelvin Kai-Wang To / Honglin Chen / Kwok-Yung Yuen / Shangyu Dang / Zhiwei Chen /  Abstract: The strikingly high transmissibility and antibody evasion of SARS-CoV-2 Omicron variants have posed great challenges to the efficacy of current vaccines and antibody immunotherapy. Here, we screen 34 ...The strikingly high transmissibility and antibody evasion of SARS-CoV-2 Omicron variants have posed great challenges to the efficacy of current vaccines and antibody immunotherapy. Here, we screen 34 BNT162b2-vaccinees and isolate a public broadly neutralizing antibody ZCB11 derived from the IGHV1-58 family. ZCB11 targets viral receptor-binding domain specifically and neutralizes all SARS-CoV-2 variants of concern, especially with great potency against authentic Omicron and Delta variants. Pseudovirus-based mapping of 57 naturally occurred spike mutations or deletions reveals that S371L results in 11-fold neutralization resistance, but it is rescued by compensating mutations in Omicron variants. Cryo-EM analysis demonstrates that ZCB11 heavy chain predominantly interacts with Omicron spike trimer with receptor-binding domain in up conformation blocking ACE2 binding. In addition, prophylactic or therapeutic ZCB11 administration protects lung infection against Omicron viral challenge in golden Syrian hamsters. These results suggest that vaccine-induced ZCB11 is a promising broadly neutralizing antibody for biomedical interventions against pandemic SARS-CoV-2. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33195.map.gz emd_33195.map.gz | 62.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33195-v30.xml emd-33195-v30.xml emd-33195.xml emd-33195.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33195.png emd_33195.png | 57.1 KB | ||
| Others |  emd_33195_half_map_1.map.gz emd_33195_half_map_1.map.gz emd_33195_half_map_2.map.gz emd_33195_half_map_2.map.gz | 116.2 MB 116.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33195 http://ftp.pdbj.org/pub/emdb/structures/EMD-33195 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33195 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33195 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33195.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33195.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
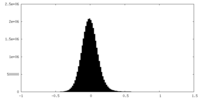
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_33195_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
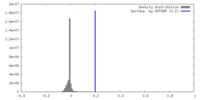
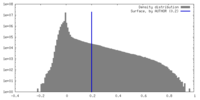
| Density Histograms |
-Half map: #2
| File | emd_33195_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : The complex of ZCB11 against Fab SARS-CoV-2 Omicron Spike
| Entire | Name: The complex of ZCB11 against Fab SARS-CoV-2 Omicron Spike |
|---|---|
| Components |
|
-Supramolecule #1: The complex of ZCB11 against Fab SARS-CoV-2 Omicron Spike
| Supramolecule | Name: The complex of ZCB11 against Fab SARS-CoV-2 Omicron Spike type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: the formation of Spike-Fab complex, the Omicron Spike trimer (2.2mg/ml) was mixed with Fab (2.9mg/ml) in 1x PBS buffer at a molar ratio of 1:1.2 (Spike monomer:Fab) and incubated on ice for 40 minutes. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #2: The ZCB11 fab
| Supramolecule | Name: The ZCB11 fab / type: complex / Chimera: Yes / ID: 2 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: The spike protein of SARS-CoV-2 Omicron variant
| Supramolecule | Name: The spike protein of SARS-CoV-2 Omicron variant / type: complex / Chimera: Yes / ID: 3 / Parent: 1 / Macromolecule list: #1 |
|---|
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average exposure time: 4.5 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Software - Name: cryoSPARC (ver. 2.15) |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.56 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 2.15) / Number images used: 86891 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2.15) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2.15) |
 Movie
Movie Controller
Controller


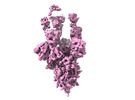



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)