[English] 日本語
 Yorodumi
Yorodumi- EMDB-33188: Complex map of Clostridioides difficile enzymatic component (CDTa... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Complex map of Clostridioides difficile enzymatic component (CDTa) and binding component (CDTb) di-heptamer | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / Translocation / Oligomer / Unfoldase / TOXIN | |||||||||
| Biological species |  Clostridioides difficile (bacteria) Clostridioides difficile (bacteria) | |||||||||
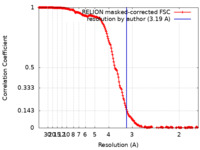
| Method | single particle reconstruction / cryo EM / Resolution: 3.19 Å | |||||||||
 Authors Authors | Yamada T / Kawamoto A / Yoshida T / Sato Y / Kato T / Tsuge H | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Cryo-EM structures of the translocational binary toxin complex CDTa-bound CDTb-pore from Clostridioides difficile. Authors: Akihiro Kawamoto / Tomohito Yamada / Toru Yoshida / Yusui Sato / Takayuki Kato / Hideaki Tsuge /  Abstract: Some bacteria express a binary toxin translocation system, consisting of an enzymatic subunit and translocation pore, that delivers enzymes into host cells through endocytosis. The most clinically ...Some bacteria express a binary toxin translocation system, consisting of an enzymatic subunit and translocation pore, that delivers enzymes into host cells through endocytosis. The most clinically important bacterium with such a system is Clostridioides difficile (formerly Clostridium). The CDTa and CDTb proteins from its system represent important therapeutic targets. CDTb has been proposed to be a di-heptamer, but its physiological heptameric structure has not yet been reported. Here, we report the cryo-EM structure of CDTa bound to the CDTb-pore, which reveals that CDTa binding induces partial unfolding and tilting of the first CDTa α-helix. In the CDTb-pore, an NSS-loop exists in 'in' and 'out' conformations, suggesting its involvement in substrate translocation. Finally, 3D variability analysis revealed CDTa movements from a folded to an unfolded state. These dynamic structural information provide insights into drug design against hypervirulent C. difficile strains. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33188.map.gz emd_33188.map.gz | 48.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33188-v30.xml emd-33188-v30.xml emd-33188.xml emd-33188.xml | 15.6 KB 15.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33188_fsc.xml emd_33188_fsc.xml | 18.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_33188.png emd_33188.png | 104.2 KB | ||
| Others |  emd_33188_half_map_1.map.gz emd_33188_half_map_1.map.gz emd_33188_half_map_2.map.gz emd_33188_half_map_2.map.gz | 391.4 MB 391.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33188 http://ftp.pdbj.org/pub/emdb/structures/EMD-33188 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33188 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33188 | HTTPS FTP |
-Validation report
| Summary document |  emd_33188_validation.pdf.gz emd_33188_validation.pdf.gz | 905.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33188_full_validation.pdf.gz emd_33188_full_validation.pdf.gz | 905.2 KB | Display | |
| Data in XML |  emd_33188_validation.xml.gz emd_33188_validation.xml.gz | 25.7 KB | Display | |
| Data in CIF |  emd_33188_validation.cif.gz emd_33188_validation.cif.gz | 34.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33188 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33188 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33188 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33188 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33188.map.gz / Format: CCP4 / Size: 488.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33188.map.gz / Format: CCP4 / Size: 488.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_33188_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
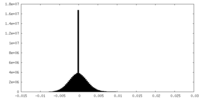
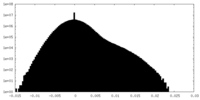
| Density Histograms |
-Half map: #2
| File | emd_33188_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
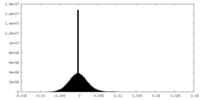
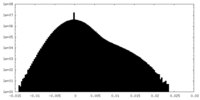
| Density Histograms |
- Sample components
Sample components
-Entire : CDTa-bound CDTb-pore (long)
| Entire | Name: CDTa-bound CDTb-pore (long) |
|---|---|
| Components |
|
-Supramolecule #1: CDTa-bound CDTb-pore (long)
| Supramolecule | Name: CDTa-bound CDTb-pore (long) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Clostridioides difficile (bacteria) Clostridioides difficile (bacteria) |
-Supramolecule #2: CDTb di-heptamer
| Supramolecule | Name: CDTb di-heptamer / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Clostridioides difficile (bacteria) Clostridioides difficile (bacteria) |
-Supramolecule #3: CdtA
| Supramolecule | Name: CdtA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.58 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 11284 / Average exposure time: 3.36 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z
Z Y
Y X
X