[English] 日本語
 Yorodumi
Yorodumi- EMDB-32678: Cryo-EM structure of SARS-CoV-2 recombinant spike protein STFK162... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
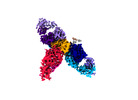
| Title | Cryo-EM structure of SARS-CoV-2 recombinant spike protein STFK1628x in complex with three neutralizing antibodies | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / spike / vaccine / neutralizing antibody / Cryo-EM / VIRAL PROTEIN / IMMUNE SYSTEM-VIRAL PROTEIN complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationMaturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell ...Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / membrane fusion / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / receptor ligand activity / symbiont-mediated suppression of host innate immune response / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.88 Å | |||||||||
 Authors Authors | Zheng Q / Sun H | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cell Host Microbe / Year: 2022 Journal: Cell Host Microbe / Year: 2022Title: Lineage-mosaic and mutation-patched spike proteins for broad-spectrum COVID-19 vaccine. Authors: Yangtao Wu / Shaojuan Wang / Yali Zhang / Lunzhi Yuan / Qingbing Zheng / Min Wei / Yang Shi / Zikang Wang / Jian Ma / Kai Wang / Meifeng Nie / Jin Xiao / Zehong Huang / Peiwen Chen / Huilin ...Authors: Yangtao Wu / Shaojuan Wang / Yali Zhang / Lunzhi Yuan / Qingbing Zheng / Min Wei / Yang Shi / Zikang Wang / Jian Ma / Kai Wang / Meifeng Nie / Jin Xiao / Zehong Huang / Peiwen Chen / Huilin Guo / Miaolin Lan / Jingjing Xu / Wangheng Hou / Yunda Hong / Dabing Chen / Hui Sun / Hualong Xiong / Ming Zhou / Che Liu / Wenjie Guo / Huiyu Guo / Jiahua Gao / Congling Gan / Zhixiong Li / Haitao Zhang / Xinrui Wang / Shaowei Li / Tong Cheng / Qinjian Zhao / Yixin Chen / Ting Wu / Tianying Zhang / Jun Zhang / Hua Cao / Huachen Zhu / Quan Yuan / Yi Guan / Ningshao Xia /  Abstract: SARS-CoV-2 spread in humans results in continuous emergence of new variants, highlighting the need for vaccines with broad-spectrum antigenic coverage. Using inter-lineage chimera and mutation-patch ...SARS-CoV-2 spread in humans results in continuous emergence of new variants, highlighting the need for vaccines with broad-spectrum antigenic coverage. Using inter-lineage chimera and mutation-patch strategies, we engineered a recombinant monomeric spike variant (STFK1628x) that contains key regions and residues across multiple SAR-CoV-2 variants. STFK1628x demonstrated high immunogenicity and mutually complementary antigenicity to its prototypic form (STFK). In hamsters, a bivalent vaccine composed of STFK and STFK1628x elicited high titers of broad-spectrum neutralizing antibodies to 19 circulating SARS-CoV-2 variants, including Omicron sublineages BA.1, BA.1.1, BA.2, BA.2.12.1, BA.2.75, and BA.4/5. Furthermore, this vaccine conferred robust protection against intranasal challenges by either SARS-CoV-2 ancestral strain or immune-evasive Beta and Omicron BA.1. Strikingly, vaccination with the bivalent vaccine in hamsters effectively blocked within-cage virus transmission of ancestral SARS-CoV-2, Beta variant, and Omicron BA.1 to unvaccinated sentinels. Thus, our study provided insight and antigen candidates for the development of next-generation COVID-19 vaccines. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32678.map.gz emd_32678.map.gz | 230.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32678-v30.xml emd-32678-v30.xml emd-32678.xml emd-32678.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_32678.png emd_32678.png | 39.8 KB | ||
| Filedesc metadata |  emd-32678.cif.gz emd-32678.cif.gz | 6.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32678 http://ftp.pdbj.org/pub/emdb/structures/EMD-32678 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32678 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32678 | HTTPS FTP |
-Validation report
| Summary document |  emd_32678_validation.pdf.gz emd_32678_validation.pdf.gz | 469.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_32678_full_validation.pdf.gz emd_32678_full_validation.pdf.gz | 468.8 KB | Display | |
| Data in XML |  emd_32678_validation.xml.gz emd_32678_validation.xml.gz | 7.1 KB | Display | |
| Data in CIF |  emd_32678_validation.cif.gz emd_32678_validation.cif.gz | 8.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32678 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32678 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32678 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32678 | HTTPS FTP |
-Related structure data
| Related structure data |  7wp8MC  7wp6C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32678.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32678.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.778 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : Cryo-EM structure of SARS-CoV-2 recombinant spike protein STFK162...
+Supramolecule #1: Cryo-EM structure of SARS-CoV-2 recombinant spike protein STFK162...
+Supramolecule #2: neutralizing antibodies
+Supramolecule #3: spike protein STFK1628x
+Macromolecule #1: 83H7 light chain
+Macromolecule #2: 83H7 heavy chain
+Macromolecule #3: 2B4 heavy chain
+Macromolecule #4: Spike glycoprotein
+Macromolecule #5: 2B4 light chain
+Macromolecule #6: 85F7 heavy chain
+Macromolecule #7: 85F7 light chain
+Macromolecule #9: 2-acetamido-2-deoxy-beta-D-glucopyranose
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.88 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 115589 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















