+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The state 3 of Omicron Spike with bispecific antibody FD01 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Omicron / Spike / Bispecific antibody / Viral protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / membrane fusion ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / membrane fusion / Attachment and Entry / entry receptor-mediated virion attachment to host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.91 Å | |||||||||
 Authors Authors | Zhang X / Zhan WQ / Chen ZG / Sun L | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2022 Journal: Cell Discov / Year: 2022Title: Combating the SARS-CoV-2 Omicron (BA.1) and BA.2 with potent bispecific antibodies engineered from non-Omicron neutralizing antibodies. Authors: Yingdan Wang / Xiang Zhang / Yunping Ma / Yanqun Wang / Wuqiang Zhan / Qinwen Zheng / Meng Zhang / Ping Ji / Mei Liu / Qianying Liu / Tingting Sun / Tongyu Zhu / Yumei Wen / Lei Sun / Jincun ...Authors: Yingdan Wang / Xiang Zhang / Yunping Ma / Yanqun Wang / Wuqiang Zhan / Qinwen Zheng / Meng Zhang / Ping Ji / Mei Liu / Qianying Liu / Tingting Sun / Tongyu Zhu / Yumei Wen / Lei Sun / Jincun Zhao / Fan Wu / Zhenguo Chen / Jinghe Huang /  Abstract: The highly mutated and transmissible Omicron (BA.1) and its more contagious lineage BA.2 have provoked serious concerns over their decreased sensitivity to the current COVID-19 vaccines and evasion ...The highly mutated and transmissible Omicron (BA.1) and its more contagious lineage BA.2 have provoked serious concerns over their decreased sensitivity to the current COVID-19 vaccines and evasion from most anti-SARS-CoV-2 neutralizing antibodies (NAbs). In this study, we explored the possibility of combating the Omicron and BA.2 by constructing bispecific antibodies based on non-Omicron NAbs. We engineered 10 IgG-like bispecific antibodies with non-Omicron NAbs named GW01, 16L9, 4L12, and REGN10987 by fusing the single-chain variable fragments (scFvs) of two antibodies through a linker and then connecting them to the Fc region of IgG1. Surprisingly, 8 out of 10 bispecific antibodies showed high binding affinities to the Omicron receptor-binding domain (RBD) and exhibited extreme breadth and potency against pseudotyped SARS-CoV-2 variants of concern (VOCs) including Omicron and BA.2, with geometric mean of 50% inhibitory concentration (GM IC) values ranging from 4.5 ng/mL to 103.94 ng/mL, as well as the authentic BA.1.1. Six bispecific antibodies containing the cross-NAb GW01 not only neutralized Omicron and BA.2, but also neutralized the sarbecoviruses including SARS-CoV and SARS-related coronaviruses (SARSr-CoVs) RS3367 and WIV1, with GM IC ranging from 11.6 ng/mL to 103.9 ng/mL. Mapping analyses of 42 spike (S) variant single mutants of Omicron and BA.2 elucidated that these bispecific antibodies accommodated the S371L/F mutations, which were resistant to most of the non-Omicron NAbs. A cryo-electron microscopy (cryo-EM) structure study of the representative bispecific antibody GW01-16L9 (FD01) in its native full-length IgG form in complex with the Omicron S trimer revealed 5 distinct trimers and one novel trimer dimer conformation. 16L9 scFv binds the receptor-binding motif (RBM), while GW01 scFv binds a epitope outside the RBM. Two scFvs of the bispecific antibody synergistically induced the RBD-down conformation into 3 RBD-up conformation, improved the affinity between IgG and the Omicron RBD, induced the formation of trimer dimer, and inhibited RBD binding to ACE2. The trimer dimer conformation might induce the aggregation of virions and contribute to the neutralization ability of FD01. These novel bispecific antibodies are strong candidates for the treatment and prevention of infection with the Omicron, BA.2, VOCs, and other sarbecoviruses. Engineering bispecific antibodies based on non-Omicron NAbs could turn the majority of NAbs into a powerful arsenal to aid the battle against the pandemic. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32657.map.gz emd_32657.map.gz | 106 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32657-v30.xml emd-32657-v30.xml emd-32657.xml emd-32657.xml | 16.8 KB 16.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32657_fsc.xml emd_32657_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_32657.png emd_32657.png | 60.4 KB | ||
| Filedesc metadata |  emd-32657.cif.gz emd-32657.cif.gz | 7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32657 http://ftp.pdbj.org/pub/emdb/structures/EMD-32657 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32657 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32657 | HTTPS FTP |
-Related structure data
| Related structure data |  7wosMC  7wopC  7woqC  7worC  7wouC  7wovC  7wowC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32657.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32657.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.064 Å | ||||||||||||||||||||||||||||||||||||
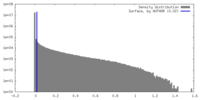
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Omicron Spike with 16L9 and GW01
| Entire | Name: Omicron Spike with 16L9 and GW01 |
|---|---|
| Components |
|
-Supramolecule #1: Omicron Spike with 16L9 and GW01
| Supramolecule | Name: Omicron Spike with 16L9 and GW01 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|
-Supramolecule #2: Omicron Spike
| Supramolecule | Name: Omicron Spike / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: 16L9 and GW01
| Supramolecule | Name: 16L9 and GW01 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 Details: residues 682-685 are substitution at furin cleavage site. T4 fibritin trimerization motif, twin strep II tag and 8-His were introduced at C-terminal. Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 142.506 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHVISG TNGTKRFDNP VLPFNDGVY FASIEKSNII RGWIFGTTLD SKTQSLLIVN NATNVVIKVC EFQFCNDPFL DHKNNKSWME SEFRVYSSAN N CTFEYVSQ ...String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHVISG TNGTKRFDNP VLPFNDGVY FASIEKSNII RGWIFGTTLD SKTQSLLIVN NATNVVIKVC EFQFCNDPFL DHKNNKSWME SEFRVYSSAN N CTFEYVSQ PFLMDLEGKQ GNFKNLREFV FKNIDGYFKI YSKHTPIIVR EPEDLPQGFS ALEPLVDLPI GINITRFQTL LA LHRSYLT PGDSSSGWTA GAAAYYVGYL QPRTFLLKYN ENGTITDAVD CALDPLSETK CTLKSFTVEK GIYQTSNFRV QPT ESIVRF PNITNLCPFD EVFNATRFAS VYAWNRKRIS NCVADYSVLY NLAPFFTFKC YGVSPTKLND LCFTNVYADS FVIR GDEVR QIAPGQTGNI ADYNYKLPDD FTGCVIAWNS NKLDSKVSGN YNYLYRLFRK SNLKPFERDI STEIYQAGNK PCNGV AGFN CYFPLRSYSF RPTYGVGHQP YRVVVLSFEL LHAPATVCGP KKSTNLVKNK CVNFNFNGLK GTGVLTESNK KFLPFQ QFG RDIADTTDAV RDPQTLEILD ITPCSFGGVS VITPGTNTSN QVAVLYQGVN CTEVPVAIHA DQLTPTWRVY STGSNVF QT RAGCLIGAEY VNNSYECDIP IGAGICASYQ TQTKSHGSAS SVASQSIIAY TMSLGAENSV AYSNNSIAIP TNFTISVT T EILPVSMTKT SVDCTMYICG DSTECSNLLL QYGSFCTQLK RALTGIAVEQ DKNTQEVFAQ VKQIYKTPPI KYFGGFNFS QILPDPSKPS KRSPIEDLLF NKVTLADAGF IKQYGDCLGD IAARDLICAQ KFKGLTVLPP LLTDEMIAQY TSALLAGTIT SGWTFGAGP ALQIPFPMQM AYRFNGIGVT QNVLYENQKL IANQFNSAIG KIQDSLSSTP SALGKLQDVV NHNAQALNTL V KQLSSKFG AISSVLNDIF SRLDPPEAEV QIDRLITGRL QSLQTYVTQQ LIRAAEIRAS ANLAATKMSE CVLGQSKRVD FC GKGYHLM SFPQSAPHGV VFLHVTYVPA QEKNFTTAPA ICHDGKAHFP REGVFVSNGT HWFVTQRNFY EPQIITTDNT FVS GNCDVV IGIVNNTVYD PLQPELDSFK EELDKYFKNH TSPDVDLGDI SGINASVVNI QKEIDRLNEV AKNLNESLID LQEL GKYEQ GSGYIPEAPR DGQAYVRKDG EWVFLSTFLS GLEVLFQGPG GWSHPQFEKG GGSGGGSGGS AWSHPQFEKG GSHHH HHHH H UniProtKB: Spike glycoprotein |
-Macromolecule #2: 16L9 Fv
| Macromolecule | Name: 16L9 Fv / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.730025 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSVLTQPPSA SGSPGQSVTI SCTGTSSDFG GYNSVSWYQQ HPGKAPKLMI YEVSKRPSGV PDRFSGSKSG NTASLTVSGL QAEDEADYY CSSYAGSNNF DVFGTGTKVT VLGGGGSGGG GSGGGGSEVQ LVESGGGLIQ PGGSLRLSCA ASGFTVSSNY M SWVRQAPG ...String: QSVLTQPPSA SGSPGQSVTI SCTGTSSDFG GYNSVSWYQQ HPGKAPKLMI YEVSKRPSGV PDRFSGSKSG NTASLTVSGL QAEDEADYY CSSYAGSNNF DVFGTGTKVT VLGGGGSGGG GSGGGGSEVQ LVESGGGLIQ PGGSLRLSCA ASGFTVSSNY M SWVRQAPG KGLEWVSVIY SGGSTYYADS VKGRFTISRD NSENTLYLQM NSLRAEDTAV YYCARGEIQP YYYYGMDVWG QG TTVTVSS |
-Macromolecule #3: GW01 Fv
| Macromolecule | Name: GW01 Fv / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.186488 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSVLTQPPSA SGTPGQRVTI SCSGSSSNIG SNTVNWYQQL PGTAPKLLIY SNNQRPSGVP DRFSGSKSGT SASLAISGLQ SEDEADYYC AAWDDSLNWV FGGGTKLTVL GGGGSGGGGS GGGGSEVQLV ESGGGVVQPG GSLRLSCAAS GFRFDDHAMH W VRQAPGKG ...String: QSVLTQPPSA SGTPGQRVTI SCSGSSSNIG SNTVNWYQQL PGTAPKLLIY SNNQRPSGVP DRFSGSKSGT SASLAISGLQ SEDEADYYC AAWDDSLNWV FGGGTKLTVL GGGGSGGGGS GGGGSEVQLV ESGGGVVQPG GSLRLSCAAS GFRFDDHAMH W VRQAPGKG LEWVSVISGD GGSTYYADSV KGRFSISRDD SKNSLYLQMN SLRTEDTALY YCAKDRSYGP PDVFNYEYGM DV WGQGTTV TVSS |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 21 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)