[English] 日本語
 Yorodumi
Yorodumi- EMDB-31176: human alpha 7 nicotinic acetylcholine receptor bound to EVP-6124 ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31176 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | human alpha 7 nicotinic acetylcholine receptor bound to EVP-6124 and PNU-120596 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | alpha 7 / nicotinic acetylcholine receptor / agonist / PAM / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsensory processing / synaptic transmission involved in micturition / response to acetylcholine / dendrite arborization / Highly calcium permeable postsynaptic nicotinic acetylcholine receptors / acetylcholine receptor activity / acetylcholine-gated channel complex / chloride channel regulator activity / regulation of amyloid fibril formation / acetylcholine-gated monoatomic cation-selective channel activity ...sensory processing / synaptic transmission involved in micturition / response to acetylcholine / dendrite arborization / Highly calcium permeable postsynaptic nicotinic acetylcholine receptors / acetylcholine receptor activity / acetylcholine-gated channel complex / chloride channel regulator activity / regulation of amyloid fibril formation / acetylcholine-gated monoatomic cation-selective channel activity / short-term memory / acetylcholine binding / dendritic spine organization / regulation of amyloid precursor protein catabolic process / acetylcholine receptor signaling pathway / positive regulation of amyloid-beta formation / negative regulation of amyloid-beta formation / response to amyloid-beta / positive regulation of protein metabolic process / negative regulation of tumor necrosis factor production / modulation of excitatory postsynaptic potential / plasma membrane raft / monoatomic ion channel activity / toxic substance binding / monoatomic ion transport / negative regulation of cytokine production involved in inflammatory response / positive regulation of long-term synaptic potentiation / positive regulation of excitatory postsynaptic potential / response to nicotine / negative regulation of canonical NF-kappaB signal transduction / excitatory postsynaptic potential / regulation of membrane potential / synapse organization / cognition / calcium channel activity / memory / positive regulation of angiogenesis / intracellular calcium ion homeostasis / calcium ion transport / transmembrane signaling receptor activity / amyloid-beta binding / monoatomic ion transmembrane transport / chemical synaptic transmission / learning or memory / response to hypoxia / positive regulation of ERK1 and ERK2 cascade / postsynaptic membrane / positive regulation of MAPK cascade / neuron projection / postsynapse / positive regulation of cell population proliferation / synapse / dendrite / endoplasmic reticulum membrane / signal transduction / protein homodimerization activity / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.02 Å | |||||||||
 Authors Authors | Liu S / Zhao Y | |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2021 Journal: Cell Res / Year: 2021Title: Structural basis of human α7 nicotinic acetylcholine receptor activation. Authors: Yue Zhao / Sanling Liu / Yingxin Zhou / Mengge Zhang / Haopeng Chen / H Eric Xu / Demeng Sun / Lei Liu / Changlin Tian /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31176.map.gz emd_31176.map.gz | 4.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31176-v30.xml emd-31176-v30.xml emd-31176.xml emd-31176.xml | 10.4 KB 10.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31176.png emd_31176.png | 121.1 KB | ||
| Filedesc metadata |  emd-31176.cif.gz emd-31176.cif.gz | 5.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31176 http://ftp.pdbj.org/pub/emdb/structures/EMD-31176 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31176 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31176 | HTTPS FTP |
-Related structure data
| Related structure data |  7ektMC  7ekiC  7ekpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31176.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31176.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.014 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
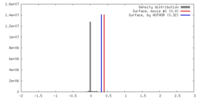
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human alpha7 nicotinic acetylcholine receptor bound to EVP-6124 a...
| Entire | Name: Human alpha7 nicotinic acetylcholine receptor bound to EVP-6124 and PNU-120596 |
|---|---|
| Components |
|
-Supramolecule #1: Human alpha7 nicotinic acetylcholine receptor bound to EVP-6124 a...
| Supramolecule | Name: Human alpha7 nicotinic acetylcholine receptor bound to EVP-6124 and PNU-120596 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Neuronal acetylcholine receptor subunit alpha-7
| Macromolecule | Name: Neuronal acetylcholine receptor subunit alpha-7 / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.505176 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MRCSPGGVWL ALAASLLHVS LQGEFQRKLY KELVKNYNPL ERPVANDSQP LTVYFSLSLL QIMDVDEKNQ VLTTNIWLQM SWTDHYLQW NVSEYPGVKT VRFPDGQIWK PDILLYNSAD ERFDATFHTN VLVNSSGHCQ YLPPGIFKSS CYIDVRWFPF D VQHCKLKF ...String: MRCSPGGVWL ALAASLLHVS LQGEFQRKLY KELVKNYNPL ERPVANDSQP LTVYFSLSLL QIMDVDEKNQ VLTTNIWLQM SWTDHYLQW NVSEYPGVKT VRFPDGQIWK PDILLYNSAD ERFDATFHTN VLVNSSGHCQ YLPPGIFKSS CYIDVRWFPF D VQHCKLKF GSWSYGGWSL DLQMQEADIS GYIPNGEWDL VGIPGKRSER FYECCKEPYP DVTFTVTMRR RTLYYGLNLL IP CVLISAL ALLVFLLPAD SGEKISLGIT VLLSLTVFML LVAEIMPATS DSVPLIAQYF ASTMIIVGLS VVVTVIVLQY HHH DPDGGK MPKWTRVILL NWCAWFLRMK RPGEDKVRPA CQHKQRRCSL ASVEMSAVAP PPASNGNLLY IGFRGLDGVH CVPT PDSGV VCGRMACSPT HDEHLLHGGQ PPEGDPDLAK ILEEVRYIAN RFRCQDESEA VCSEWKFAAC VVDRLCLMAF SVFTI ICTI GILMSAPNFV EAVSKDFA UniProtKB: Neuronal acetylcholine receptor subunit alpha-7 |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 5 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #4: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 4 / Number of copies: 5 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #5: 7-chloro-N-(quinuclidin-3-yl)benzo[b]thiophene-2-carboxamide
| Macromolecule | Name: 7-chloro-N-(quinuclidin-3-yl)benzo[b]thiophene-2-carboxamide type: ligand / ID: 5 / Number of copies: 5 / Formula: I33 |
|---|---|
| Molecular weight | Theoretical: 320.837 Da |
| Chemical component information |  ChemComp-I33: |
-Macromolecule #6: N-(5-Chloro-2,4-dimethoxyphenyl)-N'-(5-methyl-3-isoxazolyl)-urea
| Macromolecule | Name: N-(5-Chloro-2,4-dimethoxyphenyl)-N'-(5-methyl-3-isoxazolyl)-urea type: ligand / ID: 6 / Number of copies: 5 / Formula: I34 |
|---|---|
| Molecular weight | Theoretical: 311.721 Da |
| Chemical component information | 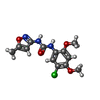 ChemComp-I34: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 62.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.02 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 46899 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)