[English] 日本語
 Yorodumi
Yorodumi- EMDB-29940: Cryo-EM Structure of the Prostaglandin E2 Receptor 4 Coupled to G... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of the Prostaglandin E2 Receptor 4 Coupled to G Protein | |||||||||
 Map data Map data | PGE2-EP4-Gi-map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of eosinophil extravasation / prostaglandin E receptor activity / negative regulation of integrin activation / Prostanoid ligand receptors / T-helper cell differentiation / regulation of stress fiber assembly / negative regulation of cytokine production / response to mechanical stimulus / JNK cascade / ERK1 and ERK2 cascade ...negative regulation of eosinophil extravasation / prostaglandin E receptor activity / negative regulation of integrin activation / Prostanoid ligand receptors / T-helper cell differentiation / regulation of stress fiber assembly / negative regulation of cytokine production / response to mechanical stimulus / JNK cascade / ERK1 and ERK2 cascade / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration / positive regulation of relaxation of smooth muscle / Adenylate cyclase inhibitory pathway / D2 dopamine receptor binding / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / cellular response to forskolin / positive regulation of cytokine production / regulation of mitotic spindle organization / chemokine-mediated signaling pathway / Regulation of insulin secretion / neuropeptide signaling pathway / response to prostaglandin E / cellular response to mechanical stimulus / positive regulation of cholesterol biosynthetic process / negative regulation of insulin secretion / bone development / G protein-coupled receptor binding / negative regulation of inflammatory response / response to peptide hormone / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G-protein beta/gamma-subunit complex binding / centriolar satellite / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / positive regulation of inflammatory response / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / GDP binding / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / G alpha (12/13) signalling events / heterotrimeric G-protein complex / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Thrombin signalling through proteinase activated receptors (PARs) / sperm principal piece / signaling receptor complex adaptor activity / positive regulation of cytosolic calcium ion concentration / retina development in camera-type eye / GTPase binding / G protein activity / Ca2+ pathway / midbody / cell cortex / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Ras protein signal transduction / Extra-nuclear estrogen signaling / cell population proliferation / immune response / ciliary basal body / G protein-coupled receptor signaling pathway / inflammatory response / cell division / lysosomal membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Huang SM / Xiong MY / Liu L / Mu J / Sheng C / Sun J | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Single hormone or synthetic agonist induces G/G coupling selectivity of EP receptors via distinct binding modes and propagating paths. Authors: Shen-Ming Huang / Meng-Yao Xiong / Lei Liu / Jianqiang Mu / Ming-Wei Wang / Ying-Li Jia / Kui Cai / Lu Tie / Chao Zhang / Sheng Cao / Xin Wen / Jia-Le Wang / Sheng-Chao Guo / Yu Li / Chang- ...Authors: Shen-Ming Huang / Meng-Yao Xiong / Lei Liu / Jianqiang Mu / Ming-Wei Wang / Ying-Li Jia / Kui Cai / Lu Tie / Chao Zhang / Sheng Cao / Xin Wen / Jia-Le Wang / Sheng-Chao Guo / Yu Li / Chang-Xiu Qu / Qing-Tao He / Bo-Yang Cai / Chenyang Xue / Shiyi Gan / Yihe Xie / Xin Cong / Zhao Yang / Wei Kong / Shuo Li / Zijian Li / Peng Xiao / Fan Yang / Xiao Yu / You-Fei Guan / Xiaoyan Zhang / Zhongmin Liu / Bao-Xue Yang / Yang Du / Jin-Peng Sun /  Abstract: To accomplish concerted physiological reactions, nature has diversified functions of a single hormone at at least two primary levels: 1) Different receptors recognize the same hormone, and 2) ...To accomplish concerted physiological reactions, nature has diversified functions of a single hormone at at least two primary levels: 1) Different receptors recognize the same hormone, and 2) different cellular effectors couple to the same hormone-receptor pair [R.P. Xiao, , re15 (2001); L. Hein, J. D. Altman, B.K. Kobilka, , 181-184 (1999); Y. Daaka, L. M. Luttrell, R. J. Lefkowitz, , 88-91 (1997)]. Not only these questions lie in the heart of hormone actions and receptor signaling but also dissecting mechanisms underlying these questions could offer therapeutic routes for refractory diseases, such as kidney injury (KI) or X-linked nephrogenic diabetes insipidus (NDI). Here, we identified that G-biased signaling, but not G activation downstream of EP4, showed beneficial effects for both KI and NDI treatments. Notably, by solving Cryo-electron microscope (cryo-EM) structures of EP3-G, EP4-G, and EP4-G in complex with endogenous prostaglandin E (PGE)or two synthetic agonists and comparing with PGE-EP2-G structures, we found that unique primary sequences of prostaglandin E2 receptor (EP) receptors and distinct conformational states of the EP4 ligand pocket govern the G/G transducer coupling selectivity through different structural propagation paths, especially via TM6 and TM7, to generate selective cytoplasmic structural features. In particular, the orientation of the PGE ω-chain and two distinct pockets encompassing agonist L902688 of EP4 were differentiated by their G/G coupling ability. Further, we identified common and distinct features of cytoplasmic side of EP receptors for G/G coupling and provide a structural basis for selective and biased agonist design of EP4 with therapeutic potential. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29940.map.gz emd_29940.map.gz | 32.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29940-v30.xml emd-29940-v30.xml emd-29940.xml emd-29940.xml | 21.7 KB 21.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_29940.png emd_29940.png | 89.6 KB | ||
| Masks |  emd_29940_msk_1.map emd_29940_msk_1.map | 38.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-29940.cif.gz emd-29940.cif.gz | 7.1 KB | ||
| Others |  emd_29940_half_map_1.map.gz emd_29940_half_map_1.map.gz emd_29940_half_map_2.map.gz emd_29940_half_map_2.map.gz | 29.6 MB 29.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29940 http://ftp.pdbj.org/pub/emdb/structures/EMD-29940 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29940 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29940 | HTTPS FTP |
-Related structure data
| Related structure data |  8gcpMC  8gcmC  8gd9C  8gdaC  8gdbC  8gdcC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29940.map.gz / Format: CCP4 / Size: 38.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29940.map.gz / Format: CCP4 / Size: 38.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PGE2-EP4-Gi-map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29940_msk_1.map emd_29940_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: PGE2-EP4-Gi-map-half1
| File | emd_29940_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PGE2-EP4-Gi-map-half1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: PGE2-EP4-Gi-map-half2
| File | emd_29940_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PGE2-EP4-Gi-map-half2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM Structure of Prostaglandin E Receptor EP4 Coupled to G Protein
| Entire | Name: Cryo-EM Structure of Prostaglandin E Receptor EP4 Coupled to G Protein |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM Structure of Prostaglandin E Receptor EP4 Coupled to G Protein
| Supramolecule | Name: Cryo-EM Structure of Prostaglandin E Receptor EP4 Coupled to G Protein type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.445059 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGAQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHESM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCS TDTKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.418086 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MHHHHHHLEV LFQGPGSSGS ELDQLRQEAE QLKNQIRDAR KACADATLSQ ITNNIDPVGR IQMRTRRTLR GHLAKIYAMH WGTDSRLLV SASQDGKLII WDSYTTNKVH AIPLRSSWVM TCAYAPSGNY VACGGLDNIC SIYNLKTREG NVRVSRELAG H TGYLSCCR ...String: MHHHHHHLEV LFQGPGSSGS ELDQLRQEAE QLKNQIRDAR KACADATLSQ ITNNIDPVGR IQMRTRRTLR GHLAKIYAMH WGTDSRLLV SASQDGKLII WDSYTTNKVH AIPLRSSWVM TCAYAPSGNY VACGGLDNIC SIYNLKTREG NVRVSRELAG H TGYLSCCR FLDDNQIVTS SGDTTCALWD IETGQQTTTF TGHTGDVMSL SLAPDTRLFV SGACDASAKL WDVREGMCRQ TF TGHESDI NAICFFPNGN AFATGSDDAT CRLFDLRADQ ELMTYSHDNI ICGITSVSFS KSGRLLLAGY DDFNCNVWDA LKA DRAGVL AGHDNRVSCL GVTDDGMAVA TGSWDSFLKI WN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein subunit gamma
| Macromolecule | Name: Guanine nucleotide-binding protein subunit gamma / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.104898 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MWRELPLGLG ELHKDHQASR KLEPELWSVS ENPPSTSMAS NNTASIAQAR KLVEQLKMEA NIDRIKVSKA AADLMAYCEA HAKEDPLLT PVPASENPFR EKKFFCAIL UniProtKB: Guanine nucleotide-binding protein subunit gamma |
-Macromolecule #4: scFv
| Macromolecule | Name: scFv / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32.898781 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MLLVNQSHQG FNKEHTSKMV SAIVLYVLLA AAAHSAFADV QLVESGGGLV QPGGSRKLSC SASGFAFSSF GMHWVRQAPE KGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSEDT AMYYCVRSIY YYGSSPFDFW GQGTTLTVSS G GGGSGGGG ...String: MLLVNQSHQG FNKEHTSKMV SAIVLYVLLA AAAHSAFADV QLVESGGGLV QPGGSRKLSC SASGFAFSSF GMHWVRQAPE KGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSEDT AMYYCVRSIY YYGSSPFDFW GQGTTLTVSS G GGGSGGGG SGGGGSDIVM TQATSSVPVT PGESVSISCR SSKSLLHSNG NTYLYWFLQR PGQSPQLLIY RMSNLASGVP DR FSGSGSG TAFTLTISRL EAEDVGVYYC MQHLEYPLTF GAGTKLELKG SLEVLFQGPA AAHHHHHHHH |
-Macromolecule #5: Prostaglandin E2 receptor EP4 subtype
| Macromolecule | Name: Prostaglandin E2 receptor EP4 subtype / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 53.173336 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MSTPGVNSSA SLSPDRLNSP VTIPAVMFIF GVVGNLVAIV VLCKSRKEQK ETTFYTLVCG LAVTDLLGTL LVSPVTIATY MKGQWPGGQ PLCEYSTFIL LFFSLSGLSI ICAMSVERYL AINHAYFYSH YVDKRLAGLT LFAVYASNVL FCALPNMGLG S SRLQYPDT ...String: MSTPGVNSSA SLSPDRLNSP VTIPAVMFIF GVVGNLVAIV VLCKSRKEQK ETTFYTLVCG LAVTDLLGTL LVSPVTIATY MKGQWPGGQ PLCEYSTFIL LFFSLSGLSI ICAMSVERYL AINHAYFYSH YVDKRLAGLT LFAVYASNVL FCALPNMGLG S SRLQYPDT WCFIDWTTNV TAHAAYSYMY AGFSSFLILA TVLCNVLVCG ALLRMHRQFM RRTSLGTEQH HAAAAASVAS RG HPAASPA LPRLSDFRRR RSFRRIAGAE IQMVILLIAT SLVVLICSIP LVVRVFVNQL YQPSLEREVS KNPDLQAIRI ASV NPILDP WIYILLRKTV LSKAIEKIKC LFCRIGGSRR ERSGQHCSDS QRTSSAMSGH SRSFISRELK EISSTSQTLL PDLS LPDLS ENGLGGRNLL PGVPGMGLAQ EDTTSLRTLR ISETSDSSQG QDSESVLLVD EAGGSGRAGP APKGSSLQVT FPSET LNLS EKCI UniProtKB: Prostaglandin E2 receptor EP4 subtype |
-Macromolecule #6: (Z)-7-[(1R,2R,3R)-3-hydroxy-2-[(E,3S)-3-hydroxyoct-1-enyl]-5-oxo-...
| Macromolecule | Name: (Z)-7-[(1R,2R,3R)-3-hydroxy-2-[(E,3S)-3-hydroxyoct-1-enyl]-5-oxo-cyclopentyl]hept-5-enoic acid type: ligand / ID: 6 / Number of copies: 1 / Formula: P2E |
|---|---|
| Molecular weight | Theoretical: 352.465 Da |
| Chemical component information | 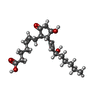 ChemComp-P2E: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source: OTHER |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 291813 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































