+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
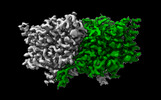
| Title | CLC-ec1 R230C/L249C/C85A at pH 4.5 100mM Cl Swap | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CLC-ec1 / ecCLC / eriC / CLC transporter / chloride proton antiporter / TRANSPORT PROTEIN | |||||||||
| Function / homology | Chloride channel, ClcA / Chloride channel, voltage gated / Chloride channel, core / Voltage gated chloride channel / voltage-gated chloride channel activity / antiporter activity / plasma membrane / H(+)/Cl(-) exchange transporter ClcA Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Fortea E / Lee S / Argyos Y / Chadda R / Ciftci D / Huysmans G / Robertson JL / Boudker O / Accardi A | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Structural basis of pH-dependent activation in a CLC transporter. Authors: Eva Fortea / Sangyun Lee / Rahul Chadda / Yiorgos Argyros / Priyanka Sandal / Robyn Mahoney-Kruszka / Hatice Didar Ciftci / Maria E Falzone / Gerard Huysmans / Janice L Robertson / Olga ...Authors: Eva Fortea / Sangyun Lee / Rahul Chadda / Yiorgos Argyros / Priyanka Sandal / Robyn Mahoney-Kruszka / Hatice Didar Ciftci / Maria E Falzone / Gerard Huysmans / Janice L Robertson / Olga Boudker / Alessio Accardi /   Abstract: CLCs are dimeric chloride channels and anion/proton exchangers that regulate processes such as muscle contraction and endo-lysosome acidification. Common gating controls their activity; its closure ...CLCs are dimeric chloride channels and anion/proton exchangers that regulate processes such as muscle contraction and endo-lysosome acidification. Common gating controls their activity; its closure simultaneously silences both protomers, and its opening allows them to independently transport ions. Mutations affecting common gating in human CLCs cause dominant genetic disorders. The structural rearrangements underlying common gating are unknown. Here, using single-particle cryo-electron microscopy, we show that the prototypical Escherichia coli CLC-ec1 undergoes large-scale rearrangements in activating conditions. The slow, pH-dependent remodeling of the dimer interface leads to the concerted opening of the intracellular H pathways and is required for transport. The more frequent formation of short water wires in the open H pathway enables Cl pore openings. Mutations at disease-causing sites favor CLC-ec1 activation and accelerate common gate opening in the human CLC-7 exchanger. We suggest that the pH activation mechanism of CLC-ec1 is related to the common gating of CLC-7. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29884.map.gz emd_29884.map.gz | 4.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29884-v30.xml emd-29884-v30.xml emd-29884.xml emd-29884.xml | 14.4 KB 14.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_29884.png emd_29884.png | 71.6 KB | ||
| Filedesc metadata |  emd-29884.cif.gz emd-29884.cif.gz | 5.4 KB | ||
| Others |  emd_29884_half_map_1.map.gz emd_29884_half_map_1.map.gz emd_29884_half_map_2.map.gz emd_29884_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29884 http://ftp.pdbj.org/pub/emdb/structures/EMD-29884 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29884 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29884 | HTTPS FTP |
-Related structure data
| Related structure data |  8ga1MC  8ga0C  8ga3C  8ga5C  8gahC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_29884.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29884.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_29884_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_29884_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of ecCLC-R230C/L249C/C85A mutant at pH 4.5 in 100mM Cl
| Entire | Name: Structure of ecCLC-R230C/L249C/C85A mutant at pH 4.5 in 100mM Cl |
|---|---|
| Components |
|
-Supramolecule #1: Structure of ecCLC-R230C/L249C/C85A mutant at pH 4.5 in 100mM Cl
| Supramolecule | Name: Structure of ecCLC-R230C/L249C/C85A mutant at pH 4.5 in 100mM Cl type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: H(+)/Cl(-) exchange transporter ClcA
| Macromolecule | Name: H(+)/Cl(-) exchange transporter ClcA / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 49.118027 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTDTPSLET PQAARLRRRQ LIRQLLERDK TPLAILFMAA VVGTLVGLAA VAFDKGVAWL QNQRMGALVH TADNYPLLLT VAFLASAVL AMFGYFLVRK YAPEAGGSGI PEIEGALEDQ RPVRWWRVLP VKFFGGLGTL GGGMVLGREG PTVQIGGNIG R MVLDIFRL ...String: MKTDTPSLET PQAARLRRRQ LIRQLLERDK TPLAILFMAA VVGTLVGLAA VAFDKGVAWL QNQRMGALVH TADNYPLLLT VAFLASAVL AMFGYFLVRK YAPEAGGSGI PEIEGALEDQ RPVRWWRVLP VKFFGGLGTL GGGMVLGREG PTVQIGGNIG R MVLDIFRL KGDEARHTLL ATGAAAGLAA AFNAPLAGIL FIIEEMRPQF RYTLISIKAV FIGVIMSTIM YCIFNHEVAL ID VGKLSDA PCNTLWLYLI LGIIFGIFGP IFNKWVLGMQ DLLHRVHGGN ITKWVLMGGA IGGLCGLLGF VAPATSGGGF NLI PIATAG NFSMGMLVFI FVARVITTLL CFSSGAPGGI FAPMLALGTV LGTAFGMVAV ELFPQYHLEA GTFAIAGMGA LLAA SIRAP LTGIILVLEM TDNYQLILPM IITGLGATLL AQFTGGKPLY SAILARTLAK QEAEQL UniProtKB: H(+)/Cl(-) exchange transporter ClcA |
-Macromolecule #2: CHLORIDE ION
| Macromolecule | Name: CHLORIDE ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CL |
|---|---|
| Molecular weight | Theoretical: 35.453 Da |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 35 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 4.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 56.11 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 718295 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)