+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | BceAB-S nucleotide free BceS state 2 | |||||||||
 Map data Map data | Final map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ABC transporter / histidine kinase / antimicrobial / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmembrane protein complex / protein histidine kinase activity / histidine kinase / phosphorelay signal transduction system / transmembrane transporter activity / transmembrane transport / response to antibiotic / ATP hydrolysis activity / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
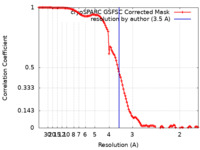
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | George NL / Orlando BJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Architecture of a complete Bce-type antimicrobial peptide resistance module. Authors: Natasha L George / Benjamin J Orlando /  Abstract: Gram-positive bacteria synthesize and secrete antimicrobial peptides that target the essential process of peptidoglycan synthesis. These antimicrobial peptides not only regulate the dynamics of ...Gram-positive bacteria synthesize and secrete antimicrobial peptides that target the essential process of peptidoglycan synthesis. These antimicrobial peptides not only regulate the dynamics of microbial communities but are also of clinical importance as exemplified by peptides such as bacitracin, vancomycin, and daptomycin. Many gram-positive species have evolved specialized antimicrobial peptide sensing and resistance machinery known as Bce modules. These modules are membrane protein complexes formed by an unusual Bce-type ABC transporter interacting with a two-component system sensor histidine kinase. In this work, we provide the first structural insight into how the membrane protein components of these modules assemble into a functional complex. A cryo-EM structure of an entire Bce module revealed an unexpected mechanism of complex assembly, and extensive structural flexibility in the sensor histidine kinase. Structures of the complex in the presence of a non-hydrolysable ATP analog reveal how nucleotide binding primes the complex for subsequent activation. Accompanying biochemical data demonstrate how the individual membrane protein components of the complex exert functional control over one another to create a tightly regulated enzymatic system. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29701.map.gz emd_29701.map.gz | 7.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29701-v30.xml emd-29701-v30.xml emd-29701.xml emd-29701.xml | 26.8 KB 26.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29701_fsc.xml emd_29701_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_29701.png emd_29701.png | 107.1 KB | ||
| Masks |  emd_29701_msk_1.map emd_29701_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-29701.cif.gz emd-29701.cif.gz | 7.7 KB | ||
| Others |  emd_29701_additional_1.map.gz emd_29701_additional_1.map.gz emd_29701_half_map_1.map.gz emd_29701_half_map_1.map.gz emd_29701_half_map_2.map.gz emd_29701_half_map_2.map.gz | 167.9 MB 165.3 MB 165.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29701 http://ftp.pdbj.org/pub/emdb/structures/EMD-29701 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29701 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29701 | HTTPS FTP |
-Related structure data
| Related structure data |  8g3lMC  8g3aC  8g3bC  8g3fC  8g4cC  8g4dC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29701.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29701.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.872 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29701_msk_1.map emd_29701_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_29701_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_29701_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_29701_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : BceAB-S
| Entire | Name: BceAB-S |
|---|---|
| Components |
|
-Supramolecule #1: BceAB-S
| Supramolecule | Name: BceAB-S / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: A membrane protein complex formed by the BceAB transporter and BceS histidine kinase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 167 KDa |
-Macromolecule #1: Bacitracin export permease protein BceB
| Macromolecule | Name: Bacitracin export permease protein BceB / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 72.262109 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNINQLILRN LKKNLRNYYL YVFALIFSVA LYFAFVTLQY DPAINEVKAS IKGAAAIKTA SILLVAVVAI FILYANTIFI KRRSKEIGL FQLIGMTKHK IFRILSAENV MLYFGSLAIG VAAGFSISKL VLMILFKIVD VKADAKLHFS EQALVQTVIV F CGIYLLIM ...String: MNINQLILRN LKKNLRNYYL YVFALIFSVA LYFAFVTLQY DPAINEVKAS IKGAAAIKTA SILLVAVVAI FILYANTIFI KRRSKEIGL FQLIGMTKHK IFRILSAENV MLYFGSLAIG VAAGFSISKL VLMILFKIVD VKADAKLHFS EQALVQTVIV F CGIYLLIM IMNYTFIKKQ SILSLFKVTS STEDKVKKIS FFQMLIGALG IVLILTGYYV SSELFGGKFK TINELFVAMS FI LGSVIIG TFLFYKGSVT FISNIIRKSK GGYLNISEVL SLSSIMFRMK SNALLLTIIT TVSALAIGLL SLAYISYYSS EKT AEQNVA ADFSFMNEKD AKLFENKLRE SNISFVKKAT PVLQANVDIA NIMDGTPKEM QGDPGNMQLA VVSDKDVKGV DVAA GEAVF SGYTDLLQKI MVFKDSGVIK VKSKHETQPL KYKGLREEFL VSYTFTSGGM PAVIVDDSLF KQLDKDKDPR IQLAQ STFI GVNVKHDDQM EKANELFQQV NKKNEHLSRL DTSAAQKSLF GMVMFIVGFL GLTFLITSGC ILYFKQMGES EDEKPS YTI LRKLGFTQGD LIKGIRIKQM YNFGIPLVVG LFHSYFAVQS GWFLFGSEVW APMIMVMVLY TALYSIFGFL SVLYYKK VI KSSL UniProtKB: Bacitracin export permease protein BceB |
-Macromolecule #2: Bacitracin export ATP-binding protein BceA
| Macromolecule | Name: Bacitracin export ATP-binding protein BceA / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 29.248377 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGHHHHHHV ILEANKIRKS YGNKLNKQEV LKGIDIHIEK GEFVSIMGAS GSGKTTLLNV LSSIDQVSHG TIHINGNDMT AMKEKQLAE FRKQHLGFIF QDYNLLDTLT VKENILLPLS ITKLSKKEAN RKFEEVAKEL GIYELRDKYP NEISGGQKQR T SAGRAFIH ...String: MSGHHHHHHV ILEANKIRKS YGNKLNKQEV LKGIDIHIEK GEFVSIMGAS GSGKTTLLNV LSSIDQVSHG TIHINGNDMT AMKEKQLAE FRKQHLGFIF QDYNLLDTLT VKENILLPLS ITKLSKKEAN RKFEEVAKEL GIYELRDKYP NEISGGQKQR T SAGRAFIH DPSIIFADEP TGALDSKSAS DLLNKLSQLN QKRNATIIMV THDPVAASYC GRVIFIKDGQ MYTQLNKGGQ DR QTFFQDI MKTQGVLGGV QHEH UniProtKB: Bacitracin export ATP-binding protein BceA |
-Macromolecule #3: Sensor protein BceS
| Macromolecule | Name: Sensor protein BceS / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO / EC number: histidine kinase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.811898 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIKAFLIERR SWIAAFLFQQ ALMLFIAFVD PSISFGNVLY MVYLCILFFI IFLWFRYRKE TAFYKSLKTW ENNLDVTAIN EPETPFEAM VERSIAGQTE HLKQTAARHR LALENEKDEL MAWIHEVKTP LTAMHLIIDR MEEKALKSQL SYEWLRIHLL L DQQLHQKR ...String: MIKAFLIERR SWIAAFLFQQ ALMLFIAFVD PSISFGNVLY MVYLCILFFI IFLWFRYRKE TAFYKSLKTW ENNLDVTAIN EPETPFEAM VERSIAGQTE HLKQTAARHR LALENEKDEL MAWIHEVKTP LTAMHLIIDR MEEKALKSQL SYEWLRIHLL L DQQLHQKR ISFIENDLSV EFIQLQPLIF KEIKDLQSWC IQKGIGFDIQ LEAKEVLSDA KWLAFIIRQL LTNAVKYSEA SE IEIKSFQ KGEQTQLQVK DCGRGIDPKD VPRIFDKGFT STTDHHDQAS TGMGLYLAKK AAAPLLIHID VESEFGAGTV FTL TFPIRN QFEHVISV UniProtKB: Sensor protein BceS |
-Macromolecule #4: PALMITIC ACID
| Macromolecule | Name: PALMITIC ACID / type: ligand / ID: 4 / Number of copies: 1 / Formula: PLM |
|---|---|
| Molecular weight | Theoretical: 256.424 Da |
| Chemical component information |  ChemComp-PLM: |
-Macromolecule #5: [(2~{R})-1-[2-azanylethoxy(oxidanyl)phosphoryl]oxy-3-hexadecanoyl...
| Macromolecule | Name: [(2~{R})-1-[2-azanylethoxy(oxidanyl)phosphoryl]oxy-3-hexadecanoyloxy-propan-2-yl] (~{Z})-octadec-9-enoate type: ligand / ID: 5 / Number of copies: 2 / Formula: 6OU |
|---|---|
| Molecular weight | Theoretical: 717.996 Da |
| Chemical component information |  ChemComp-6OU: |
-Macromolecule #6: OLEIC ACID
| Macromolecule | Name: OLEIC ACID / type: ligand / ID: 6 / Number of copies: 1 / Formula: OLA |
|---|---|
| Molecular weight | Theoretical: 282.461 Da |
| Chemical component information |  ChemComp-OLA: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: 150mM NaCl, 25mM Tris-HCl, 0.005% LMNG | ||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)