+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cas9 with double-stranded guide RNA and target DNA | |||||||||
 Map data Map data | DeepEMhancer HiRes map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cas9 / RNA / DNA / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationmaintenance of CRISPR repeat elements / 3'-5' exonuclease activity / DNA endonuclease activity / defense response to virus / Hydrolases; Acting on ester bonds / DNA binding / RNA binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Streptococcus pyogenes (bacteria) / Streptococcus pyogenes (bacteria) /  Homo sapiens (human) Homo sapiens (human) | |||||||||
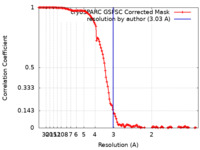
| Method | single particle reconstruction / cryo EM / Resolution: 3.03 Å | |||||||||
 Authors Authors | Korolev S / Gagnon K | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: SpCas9 with dual-guide RNA and target DNA Authors: Korolev S / Gagnon K | |||||||||
| History |
|
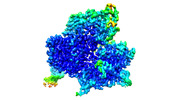
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29639.map.gz emd_29639.map.gz | 173.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29639-v30.xml emd-29639-v30.xml emd-29639.xml emd-29639.xml | 19 KB 19 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29639_fsc.xml emd_29639_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_29639.png emd_29639.png | 66.8 KB | ||
| Masks |  emd_29639_msk_1.map emd_29639_msk_1.map | 209.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-29639.cif.gz emd-29639.cif.gz | 7.1 KB | ||
| Others |  emd_29639_half_map_1.map.gz emd_29639_half_map_1.map.gz emd_29639_half_map_2.map.gz emd_29639_half_map_2.map.gz | 194.4 MB 194.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29639 http://ftp.pdbj.org/pub/emdb/structures/EMD-29639 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29639 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29639 | HTTPS FTP |
-Related structure data
| Related structure data |  8fztMC  9b2kC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29639.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29639.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DeepEMhancer HiRes map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.7 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29639_msk_1.map emd_29639_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_29639_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_29639_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CRISPR-associated endonuclease Cas9 complex with double stranded ...
| Entire | Name: CRISPR-associated endonuclease Cas9 complex with double stranded guide RNA and target DNA |
|---|---|
| Components |
|
-Supramolecule #1: CRISPR-associated endonuclease Cas9 complex with double stranded ...
| Supramolecule | Name: CRISPR-associated endonuclease Cas9 complex with double stranded guide RNA and target DNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Streptococcus pyogenes (bacteria) Streptococcus pyogenes (bacteria) |
-Macromolecule #1: CRISPR-associated endonuclease Cas9/Csn1
| Macromolecule | Name: CRISPR-associated endonuclease Cas9/Csn1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: Hydrolases; Acting on ester bonds |
|---|---|
| Source (natural) | Organism:  Streptococcus pyogenes (bacteria) Streptococcus pyogenes (bacteria) |
| Molecular weight | Theoretical: 160.774281 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDKKYSIGLD IGTNSVGWAV ITDEYKVPSK KFKVLGNTDR HSIKKNLIGA LLFDSGETAE ATRLKRTARR RYTRRKNRIC YLQEIFSNE MAKVDDSFFH RLEESFLVEE DKKHERHPIF GNIVDEVAYH EKYPTIYHLR KKLVDSTDKA DLRLIYLALA H MIKFRGHF ...String: MDKKYSIGLD IGTNSVGWAV ITDEYKVPSK KFKVLGNTDR HSIKKNLIGA LLFDSGETAE ATRLKRTARR RYTRRKNRIC YLQEIFSNE MAKVDDSFFH RLEESFLVEE DKKHERHPIF GNIVDEVAYH EKYPTIYHLR KKLVDSTDKA DLRLIYLALA H MIKFRGHF LIEGDLNPDN SDVDKLFIQL VQTYNQLFEE NPINASGVDA KAILSARLSK SRRLENLIAQ LPGEKKNGLF GN LIALSLG LTPNFKSNFD LAEDAKLQLS KDTYDDDLDN LLAQIGDQYA DLFLAAKNLS DAILLSDILR VNTEITKAPL SAS MIKRYD EHHQDLTLLK ALVRQQLPEK YKEIFFDQSK NGYAGYIDGG ASQEEFYKFI KPILEKMDGT EELLVKLNRE DLLR KQRTF DNGSIPHQIH LGELHAILRR QEDFYPFLKD NREKIEKILT FRIPYYVGPL ARGNSRFAWM TRKSEETITP WNFEE VVDK GASAQSFIER MTNFDKNLPN EKVLPKHSLL YEYFTVYNEL TKVKYVTEGM RKPAFLSGEQ KKAIVDLLFK TNRKVT VKQ LKEDYFKKIE CFDSVEISGV EDRFNASLGT YHDLLKIIKD KDFLDNEENE DILEDIVLTL TLFEDREMIE ERLKTYA HL FDDKVMKQLK RRRYTGWGRL SRKLINGIRD KQSGKTILDF LKSDGFANRN FMQLIHDDSL TFKEDIQKAQ VSGQGDSL H EHIANLAGSP AIKKGILQTV KVVDELVKVM GRHKPENIVI EMARENQTTQ KGQKNSRERM KRIEEGIKEL GSQILKEHP VENTQLQNEK LYLYYLQNGR DMYVDQELDI NRLSDYDVDH IVPQSFLKDD SIDNKVLTRS DKNRGKSDNV PSEEVVKKMK NYWRQLLNA KLITQRKFDN LTKAERGGLS ELDKAGFIKR QLVETRQITK HVAQILDSRM NTKYDENDKL IREVKVITLK S KLVSDFRK DFQFYKVREI NNYHHAHDAY LNAVVGTALI KKYPKLESEF VYGDYKVYDV RKMIAKSEQE IGKATAKYFF YS NIMNFFK TEITLANGEI RKRPLIETNG ETGEIVWDKG RDFATVRKVL SMPQVNIVKK TEVQTGGFSK ESILPKRNSD KLI ARKKDW DPKKYGGFDS PTVAYSVLVV AKVEKGKSKK LKSVKELLGI TIMERSSFEK NPIDFLEAKG YKEVKKDLII KLPK YSLFE LENGRKRMLA SAGELQKGNE LALPSKYVNF LYLASHYEKL KGSPEDNEQK QLFVEQHKHY LDEIIEQISE FSKRV ILAD ANLDKVLSAY NKHRDKPIRE QAENIIHLFT LTNLGAPAAF KYFDTTIDRK RYTSTKEVLD ATLIHQSITG LYETRI DLS QLGGDPKKKR KVMDKHHHHH H UniProtKB: CRISPR-associated endonuclease Cas9/Csn1 |
-Macromolecule #2: CrE2
| Macromolecule | Name: CrE2 / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Streptococcus pyogenes (bacteria) Streptococcus pyogenes (bacteria) |
| Molecular weight | Theoretical: 13.370906 KDa |
| Sequence | String: UACCAGCAAA ACACUCCGAU GUUUUAGAGC UAUGCUGUUU UG |
-Macromolecule #3: tracrRNA
| Macromolecule | Name: tracrRNA / type: rna / ID: 3 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Streptococcus pyogenes (bacteria) Streptococcus pyogenes (bacteria) |
| Molecular weight | Theoretical: 22.574494 KDa |
| Sequence | String: AAACAGCAUA GCAAGUUAAA AUAAGGCUAG UCCGUUAUCA ACUUGAAAAA GUGGCACCGA GUCGGUGCUU GENBANK: GENBANK: LS483330.1 |
-Macromolecule #4: E2_TS
| Macromolecule | Name: E2_TS / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.334909 KDa |
| Sequence | String: (DC)(DT)(DA)(DA)(DT)(DC)(DG)(DC)(DC)(DA) (DA)(DT)(DC)(DG)(DG)(DA)(DG)(DT)(DG)(DT) (DT)(DT)(DT)(DG)(DC)(DT)(DG)(DG)(DT) (DA)(DT)(DA)(DC)(DG)(DC)(DA)(DC)(DT)(DG) (DG) |
-Macromolecule #5: E2-NTS
| Macromolecule | Name: E2-NTS / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.290917 KDa |
| Sequence | String: (DC)(DC)(DA)(DG)(DT)(DG)(DC)(DG)(DT)(DA) (DT)(DA)(DC)(DC)(DA)(DG)(DC)(DA)(DA)(DA) (DA)(DC)(DA)(DC)(DT)(DC)(DC)(DG)(DA) (DT)(DT)(DG)(DG)(DC)(DG)(DA)(DT)(DT)(DA) (DG) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.4 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Chromatic aberration corrector: CEOS manufactured Cc corrector |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 11890 / Average exposure time: 4.28 sec. / Average electron dose: 51.37 e/Å2 / Details: Movies were collected in counted-mode |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8fzt: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)