[English] 日本語
 Yorodumi
Yorodumi- EMDB-28854: Bovine multidrug resistance protein 1 (MRP1) bound to cyclic pept... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bovine multidrug resistance protein 1 (MRP1) bound to cyclic peptide inhibitor 1 (CPI1) | |||||||||
 Map data Map data | Full map of the 3.27 angstrom density map generated from image processing in RELION | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationSphingolipid de novo biosynthesis / Heme degradation / Synthesis of Leukotrienes (LT) and Eoxins (EX) / Transport of RCbl within the body / cyclic nucleotide transport / Paracetamol ADME / ABC-family protein mediated transport / glutathione transmembrane transport / leukotriene transport / Cytoprotection by HMOX1 ...Sphingolipid de novo biosynthesis / Heme degradation / Synthesis of Leukotrienes (LT) and Eoxins (EX) / Transport of RCbl within the body / cyclic nucleotide transport / Paracetamol ADME / ABC-family protein mediated transport / glutathione transmembrane transport / leukotriene transport / Cytoprotection by HMOX1 / glutathione transmembrane transporter activity / ABC-type glutathione-S-conjugate transporter / carboxylic acid transmembrane transporter activity / ABC-type glutathione S-conjugate transporter activity / ABC-type xenobiotic transporter / ABC-type xenobiotic transporter activity / lipid transport / xenobiotic transmembrane transporter activity / xenobiotic transport / ABC-type transporter activity / positive regulation of inflammatory response / basolateral plasma membrane / response to xenobiotic stimulus / ATP hydrolysis activity / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
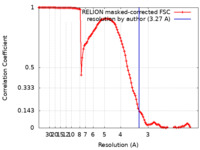
| Method | single particle reconstruction / cryo EM / Resolution: 3.27 Å | |||||||||
 Authors Authors | Pietz HL / Chen J | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: A macrocyclic peptide inhibitor traps MRP1 in a catalytically incompetent conformation. Authors: Harlan L Pietz / Ata Abbas / Zachary Lee Johnson / Michael L Oldham / Hiroaki Suga / Jue Chen /   Abstract: Adenosine triphosphate-binding cassette (ABC) transporters, such as multidrug resistance protein 1 (MRP1), protect against cellular toxicity by exporting xenobiotic compounds across the plasma ...Adenosine triphosphate-binding cassette (ABC) transporters, such as multidrug resistance protein 1 (MRP1), protect against cellular toxicity by exporting xenobiotic compounds across the plasma membrane. However, constitutive MRP1 function hinders drug delivery across the blood-brain barrier, and MRP1 overexpression in certain cancers leads to acquired multidrug resistance and chemotherapy failure. Small-molecule inhibitors have the potential to block substrate transport, but few show specificity for MRP1. Here we identify a macrocyclic peptide, named CPI1, which inhibits MRP1 with nanomolar potency but shows minimal inhibition of a related multidrug transporter P-glycoprotein. A cryoelectron microscopy (cryo-EM) structure at 3.27 Å resolution shows that CPI1 binds MRP1 at the same location as the physiological substrate leukotriene C4 (LTC). Residues that interact with both ligands contain large, flexible sidechains that can form a variety of interactions, revealing how MRP1 recognizes multiple structurally unrelated molecules. CPI1 binding prevents the conformational changes necessary for adenosine triphosphate (ATP) hydrolysis and substrate transport, suggesting it may have potential as a therapeutic candidate. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28854.map.gz emd_28854.map.gz | 117 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28854-v30.xml emd-28854-v30.xml emd-28854.xml emd-28854.xml | 25.5 KB 25.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28854_fsc.xml emd_28854_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_28854.png emd_28854.png | 126.4 KB | ||
| Masks |  emd_28854_msk_1.map emd_28854_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-28854.cif.gz emd-28854.cif.gz | 6.7 KB | ||
| Others |  emd_28854_additional_1.map.gz emd_28854_additional_1.map.gz emd_28854_additional_2.map.gz emd_28854_additional_2.map.gz emd_28854_additional_3.map.gz emd_28854_additional_3.map.gz emd_28854_additional_4.map.gz emd_28854_additional_4.map.gz emd_28854_half_map_1.map.gz emd_28854_half_map_1.map.gz emd_28854_half_map_2.map.gz emd_28854_half_map_2.map.gz | 4.7 MB 4.7 MB 4.7 MB 6.8 MB 98.3 MB 98.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28854 http://ftp.pdbj.org/pub/emdb/structures/EMD-28854 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28854 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28854 | HTTPS FTP |
-Related structure data
| Related structure data |  8f4bMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28854.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28854.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map of the 3.27 angstrom density map generated from image processing in RELION | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28854_msk_1.map emd_28854_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Full density-modified map from the 3.27 angstrom map...
| File | emd_28854_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full density-modified map from the 3.27 angstrom map generated from image processing in RELION | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: First half map of density-modified map from 3.27...
| File | emd_28854_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | First half map of density-modified map from 3.27 angstrom map generated from image processing in RELION | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Second half map of density-modified map from 3.27...
| File | emd_28854_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Second half map of density-modified map from 3.27 angstrom map generated from image processing in RELION | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Masked full map of the 3.27 angstrom density...
| File | emd_28854_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Masked full map of the 3.27 angstrom density map generated from image processing in RELION | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Second half map of the 3.27 angstrom density...
| File | emd_28854_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Second half map of the 3.27 angstrom density map generated from image processing in RELION | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: First half map of the 3.27 angstrom density...
| File | emd_28854_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | First half map of the 3.27 angstrom density map generated from image processing in RELION | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Bovine multidrug resistance protein 1 (MRP1) in complex with cycl...
| Entire | Name: Bovine multidrug resistance protein 1 (MRP1) in complex with cyclic peptide inhibitor 1 (CPI1) |
|---|---|
| Components |
|
-Supramolecule #1: Bovine multidrug resistance protein 1 (MRP1) in complex with cycl...
| Supramolecule | Name: Bovine multidrug resistance protein 1 (MRP1) in complex with cyclic peptide inhibitor 1 (CPI1) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Multidrug resistance-associated protein 1
| Macromolecule | Name: Multidrug resistance-associated protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: ABC-type xenobiotic transporter |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 148.045859 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: PNPCPESSAS FLSRITFWWI TGMMVQGYRQ PLESTDLWSL NKEDTSEQVV PVLVKNWKKE CAKSRKQPVK IVYSSKDPAK PKGSSKVDV NEEAEALIVK CPQKERDPSL FKVLYKTFGP YFLMSFLFKA VHDLMMFAGP EILKLLINFV NDKKAPEWQG Y FYTALLFI ...String: PNPCPESSAS FLSRITFWWI TGMMVQGYRQ PLESTDLWSL NKEDTSEQVV PVLVKNWKKE CAKSRKQPVK IVYSSKDPAK PKGSSKVDV NEEAEALIVK CPQKERDPSL FKVLYKTFGP YFLMSFLFKA VHDLMMFAGP EILKLLINFV NDKKAPEWQG Y FYTALLFI SACLQTLVLH QYFHICFVSG MRIKTAVIGA VYRKALVITN AARKSSTVGE IVNLMSVDAQ RFMDLATYIN MI WSAPLQV ILALYLLWLN LGPSVLAGVA VMVLMVPLNA VMAMKTKTYQ VAHMKSKDNR IKLMNEILNG IKVLKLYAWE LAF KDKVLA IRQEELKVLK KSAYLAAVGT FTWVCTPFLV ALSTFAVYVT VDENNILDAQ KAFVSLALFN ILRFPLNILP MVIS SIVQA SVSLKRLRVF LSHEDLDPDS IQRRPIKDAG ATNSITVKNA TFTWARNDPP TLHGITFSVP EGSLVAVVGQ VGCGK SSLL SALLAEMDKV EGHVTVKGSV AYVPQQAWIQ NISLRENILF GRQLQERYYK AVVEACALLP DLEILPSGDR TEIGEK GVN LSGGQKQRVS LARAVYCDSD VYLLDDPLSA VDAHVGKHIF ENVIGPKGLL KNKTRLLVTH AISYLPQMDV IIVMSGG KI SEMGSYQELL ARDGAFAEFL RTYASAEQEQ GQPEDGLAGV GGPGKEVKQM ENGMLVTDTA GKQMQRQLSS SSSYSRDV S QHHTSTAELR KPGPTEETWK LVEADKAQTG QVKLSVYWDY MKAIGLFISF LSIFLFLCNH VASLVSNYWL SLWTDDPIV NGTQEHTQVR LSVYGALGIS QGITVFGYSM AVSIGGIFAS RRLHLDLLHN VLRSPISFFE RTPSGNLVNR FSKELDTVDS MIPQVIKMF MGSLFNVIGA CIIILLATPM AAVIIPPLGL IYFFVQRFYV ASSRQLKRLE SVSRSPVYSH FNETLLGVSV I RAFEEQER FIRQSDLKVD ENQKAYYPSI VANRWLAVRL ECVGNCIVLF ASLFAVISRH SLSAGLVGLS VSYSLQVTTY LN WLVRMSS EMETNIVAVE RLKEYSETEK EAPWQIQDMA PPKDWPQVGR VEFRDYGLRY REDLDLVLKH INVTIDGGEK VGI VGRTGA GKSSLTLGLF RIKESAEGEI IIDDINIAKI GLHDLRFKIT IIPQDPVLFS GSLRMNLDPF SQYSDEEVWT SLEL AHLKG FVSALPDKLN HECAEGGENL SVGQRQLVCL ARALLRKTKI LVLDEATAAV DLETDDLIQS TIRTQFDDCT VLTIA HRLN TIMDYTRVIV LDKGEIQEWG SPSDLLQQRG LFYSMAKDSG LV UniProtKB: Multidrug resistance-associated protein 1 |
-Macromolecule #2: Cyclic peptide inhibitor 1 (CPI1)
| Macromolecule | Name: Cyclic peptide inhibitor 1 (CPI1) / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 2.286456 KDa |
| Recombinant expression | Organism: synthetic construct (others) |
| Sequence | String: (DTY)FWGNLHWYY EQFDSTC(ACE) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-50 / Number grids imaged: 1 / Number real images: 3708 / Average exposure time: 0.2 sec. / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.7000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-8f4b: |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)