+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | EsN-dhsU36mm1 local map (map 2) | |||||||||
 Map data Map data | Raw map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | promoter-bound / initiation / TRANSCRIPTION | |||||||||
| Biological species |    Aquifex aeolicus (bacteria) Aquifex aeolicus (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Mueller AU / Chen J / Darst SA | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: A general mechanism for transcription bubble nucleation in bacteria. Authors: Andreas U Mueller / James Chen / Mengyu Wu / Courtney Chiu / B Tracy Nixon / Elizabeth A Campbell / Seth A Darst /  Abstract: Bacterial transcription initiation requires σ factors for nucleation of the transcription bubble. The canonical housekeeping σ factor, σ, nucleates DNA melting via recognition of conserved bases ...Bacterial transcription initiation requires σ factors for nucleation of the transcription bubble. The canonical housekeeping σ factor, σ, nucleates DNA melting via recognition of conserved bases of the promoter -10 motif, which are unstacked and captured in pockets of σ. By contrast, the mechanism of transcription bubble nucleation and formation during the unrelated σ-mediated transcription initiation is poorly understood. Herein, we combine structural and biochemical approaches to establish that σ, like σ, captures a flipped, unstacked base in a pocket formed between its N-terminal region I (RI) and extra-long helix features. Strikingly, RI inserts into the nascent bubble to stabilize the nucleated bubble prior to engagement of the obligate ATPase activator. Our data suggest a general paradigm of transcription initiation that requires σ factors to nucleate an early melted intermediate prior to productive RNA synthesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28794.map.gz emd_28794.map.gz | 32.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28794-v30.xml emd-28794-v30.xml emd-28794.xml emd-28794.xml | 22.1 KB 22.1 KB | Display Display |  EMDB header EMDB header |
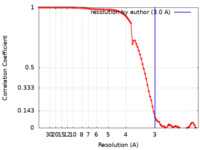
| FSC (resolution estimation) |  emd_28794_fsc.xml emd_28794_fsc.xml | 8.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_28794.png emd_28794.png | 78.1 KB | ||
| Masks |  emd_28794_msk_1.map emd_28794_msk_1.map emd_28794_msk_2.map emd_28794_msk_2.map | 64 MB 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-28794.cif.gz emd-28794.cif.gz | 4.9 KB | ||
| Others |  emd_28794_additional_1.map.gz emd_28794_additional_1.map.gz emd_28794_additional_2.map.gz emd_28794_additional_2.map.gz emd_28794_additional_3.map.gz emd_28794_additional_3.map.gz emd_28794_half_map_1.map.gz emd_28794_half_map_1.map.gz emd_28794_half_map_2.map.gz emd_28794_half_map_2.map.gz | 59.8 MB 2.4 MB 3 MB 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28794 http://ftp.pdbj.org/pub/emdb/structures/EMD-28794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28794 | HTTPS FTP |
-Validation report
| Summary document |  emd_28794_validation.pdf.gz emd_28794_validation.pdf.gz | 878.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28794_full_validation.pdf.gz emd_28794_full_validation.pdf.gz | 878.5 KB | Display | |
| Data in XML |  emd_28794_validation.xml.gz emd_28794_validation.xml.gz | 16.5 KB | Display | |
| Data in CIF |  emd_28794_validation.cif.gz emd_28794_validation.cif.gz | 21.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28794 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28794 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28794 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28794 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28794.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28794.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Raw map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.083 Å | ||||||||||||||||||||||||||||||||||||
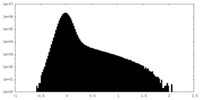
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28794_msk_1.map emd_28794_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Mask #2
| File |  emd_28794_msk_2.map emd_28794_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: B-factor sharpened map (-110.1)
| File | emd_28794_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | B-factor sharpened map (-110.1) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Locally filtered map (blocfilt)
| File | emd_28794_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally filtered map (blocfilt) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Local resolution map (blocres)
| File | emd_28794_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution map (blocres) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_28794_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_28794_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : EsN-dhsU36mm1
| Entire | Name: EsN-dhsU36mm1 |
|---|---|
| Components |
|
-Supramolecule #1: EsN-dhsU36mm1
| Supramolecule | Name: EsN-dhsU36mm1 / type: complex / ID: 1 / Parent: 0 |
|---|
-Supramolecule #2: RNA polymerase complex
| Supramolecule | Name: RNA polymerase complex / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: DNA
| Supramolecule | Name: DNA / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:   Aquifex aeolicus (bacteria) / Synthetically produced: Yes Aquifex aeolicus (bacteria) / Synthetically produced: Yes |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV Details: Octyl beta-D-glucopyranoside (beta-OG) was added to the sample to 0.1%w/v final concentration (from 10x stock) just prior to plunge vitrification. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 12294 / Average exposure time: 2.0 sec. / Average electron dose: 51.0 e/Å2 Details: dose-fractionation with 0.05 seconds per frame (=40 frames) |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)