[English] 日本語
 Yorodumi
Yorodumi- EMDB-27825: Cryo-EM structure of the endogenous core TIM23 complex from S. ce... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the endogenous core TIM23 complex from S. cerevisiae | |||||||||
 Map data Map data | Unsharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRANSLOCASE-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationTIM23 mitochondrial import inner membrane translocase complex / protein import into mitochondrial matrix / transmembrane protein transporter activity / protein-folding chaperone binding / mitochondrial inner membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
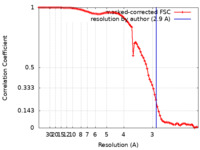
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Sim SI / Park E | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Structural basis of mitochondrial protein import by the TIM23 complex. Authors: Sue Im Sim / Yuanyuan Chen / Diane L Lynch / James C Gumbart / Eunyong Park /  Abstract: Mitochondria import nearly all of their approximately 1,000-2,000 constituent proteins from the cytosol across their double-membrane envelope. Genetic and biochemical studies have shown that the ...Mitochondria import nearly all of their approximately 1,000-2,000 constituent proteins from the cytosol across their double-membrane envelope. Genetic and biochemical studies have shown that the conserved protein translocase, termed the TIM23 complex, mediates import of presequence-containing proteins (preproteins) into the mitochondrial matrix and inner membrane. Among about ten different subunits of the TIM23 complex, the essential multipass membrane protein Tim23, together with the evolutionarily related protein Tim17, has long been postulated to form a protein-conducting channel. However, the mechanism by which these subunits form a translocation path in the membrane and enable the import process remains unclear due to a lack of structural information. Here we determined the cryo-electron microscopy structure of the core TIM23 complex (heterotrimeric Tim17-Tim23-Tim44) from Saccharomyces cerevisiae. Contrary to the prevailing model, Tim23 and Tim17 themselves do not form a water-filled channel, but instead have separate, lipid-exposed concave cavities that face in opposite directions. Our structural and biochemical analyses show that the cavity of Tim17, but not Tim23, forms the protein translocation path, whereas Tim23 probably has a structural role. The results further suggest that, during translocation of substrate polypeptides, the nonessential subunit Mgr2 seals the lateral opening of the Tim17 cavity to facilitate the translocation process. We propose a new model for the TIM23-mediated protein import and sorting mechanism, a central pathway in mitochondrial biogenesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27825.map.gz emd_27825.map.gz | 32.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27825-v30.xml emd-27825-v30.xml emd-27825.xml emd-27825.xml | 27.1 KB 27.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27825_fsc.xml emd_27825_fsc.xml | 9.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_27825.png emd_27825.png | 27.2 KB | ||
| Filedesc metadata |  emd-27825.cif.gz emd-27825.cif.gz | 7.5 KB | ||
| Others |  emd_27825_additional_1.map.gz emd_27825_additional_1.map.gz emd_27825_half_map_1.map.gz emd_27825_half_map_1.map.gz emd_27825_half_map_2.map.gz emd_27825_half_map_2.map.gz | 54.3 MB 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27825 http://ftp.pdbj.org/pub/emdb/structures/EMD-27825 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27825 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27825 | HTTPS FTP |
-Related structure data
| Related structure data |  8e1mMC  8scxC  24885  24886 M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27825.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27825.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Sharpened map
| File | emd_27825_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_27825_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_27825_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Core TIM23 complex (endogenous)
| Entire | Name: Core TIM23 complex (endogenous) |
|---|---|
| Components |
|
-Supramolecule #1: Core TIM23 complex (endogenous)
| Supramolecule | Name: Core TIM23 complex (endogenous) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Mitochondrial import inner membrane translocase subunit TIM17
| Macromolecule | Name: Mitochondrial import inner membrane translocase subunit TIM17 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.602156 KDa |
| Sequence | String: MSADHSRDPC PIVILNDFGG AFAMGAIGGV VWHGIKGFRN SPLGERGSGA MSAIKARAPV LGGNFGVWGG LFSTFDCAVK AVRKREDPW NAIIAGFFTG GALAVRGGWR HTRNSSITCA CLLGVIEGVG LMFQRYAAWQ AKPMAPPLPE APSSQPLQA UniProtKB: Mitochondrial import inner membrane translocase subunit TIM17 |
-Macromolecule #2: Mitochondrial import inner membrane translocase subunit TIM23
| Macromolecule | Name: Mitochondrial import inner membrane translocase subunit TIM23 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.266527 KDa |
| Sequence | String: MSWLFGDKTP TDDANAAVGG QDTTKPKELS LKQSLGFEPN INNIISGPGG MHVDTARLHP LAGLDKGVEY LDLEEEQLSS LEGSQGLIP SRGWTDDLCY GTGAVYLLGL GIGGFSGMMQ GLQNIPPNSP GKLQLNTVLN HITKRGPFLG NNAGILALSY N IINSTIDA ...String: MSWLFGDKTP TDDANAAVGG QDTTKPKELS LKQSLGFEPN INNIISGPGG MHVDTARLHP LAGLDKGVEY LDLEEEQLSS LEGSQGLIP SRGWTDDLCY GTGAVYLLGL GIGGFSGMMQ GLQNIPPNSP GKLQLNTVLN HITKRGPFLG NNAGILALSY N IINSTIDA LRGKHDTAGS IGAGALTGAL FKSSKGLKPM GYSSAMVAAA CAVWCSVKKR LLEK UniProtKB: Mitochondrial import inner membrane translocase subunit TIM23 |
-Macromolecule #3: Mitochondrial import inner membrane translocase subunit TIM44
| Macromolecule | Name: Mitochondrial import inner membrane translocase subunit TIM44 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 48.933418 KDa |
| Sequence | String: MHRSTFIRTS GTSSRTLTAR YRSQYTGLLV ARVLFSTSTT RAQGGNPRSP LQIFRDTFKK EWEKSQELQE NIKTLQDASG KLGESEAYK KAREAYLKAQ RGSTIVGKTL KKTGETMEHI ATKAWESELG KNTRKAAAAT AKKLDESFEP VRQTKIYKEV S EVIDDGES ...String: MHRSTFIRTS GTSSRTLTAR YRSQYTGLLV ARVLFSTSTT RAQGGNPRSP LQIFRDTFKK EWEKSQELQE NIKTLQDASG KLGESEAYK KAREAYLKAQ RGSTIVGKTL KKTGETMEHI ATKAWESELG KNTRKAAAAT AKKLDESFEP VRQTKIYKEV S EVIDDGES SRYGGFITKE QRRLKRERDL ASGKRHRAVK SNEDAGTAVV ATNIESKESF GKKVEDFKEK TVVGRSIQSL KN KLWDESE NPLIVVMRKI TNKVGGFFAE TESSRVYSQF KLMDPTFSNE SFTRHLREYI VPEILEAYVK GDVKVLKKWF SEA PFNVYA AQQKIFKEQD VYADGRILDI RGVEIVSAKL LAPQDIPVLV VGCRAQEINL YRKKKTGEIA AGDEANILMS SYAM VFTRD PEQIDDDETE GWKILEFVRG GSRQFT UniProtKB: Mitochondrial import inner membrane translocase subunit TIM44 |
-Macromolecule #4: Antibody Fab fragment light chain
| Macromolecule | Name: Antibody Fab fragment light chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.173201 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEKDTLLLWV LLLWVPGSTG DIVLTQSPAS LAVSLGQRAT ISCRASESVD IYGISFMNWF QQKPGQPPKL LIYATSNQGS GVPARFSGS GSGTDFSLNI HPMEEDDTAM YFCQQSKEVP RTFGGGTKLE IKRADAAPTV SIFPPSSEQL TSGGASVVCF L NNFYPKDI ...String: MEKDTLLLWV LLLWVPGSTG DIVLTQSPAS LAVSLGQRAT ISCRASESVD IYGISFMNWF QQKPGQPPKL LIYATSNQGS GVPARFSGS GSGTDFSLNI HPMEEDDTAM YFCQQSKEVP RTFGGGTKLE IKRADAAPTV SIFPPSSEQL TSGGASVVCF L NNFYPKDI NVKWKIDGSE RQNGVLNSWT DQDSKDSTYS MSSTLTLTKD EYERHNSYTC EATHKTSTSP IVKSFNRNEC |
-Macromolecule #5: Antibody Fab fragment heavy chain
| Macromolecule | Name: Antibody Fab fragment heavy chain / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.49499 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAVLVLLLCL VTFPSCVLSQ VQLKQSGPGL VQPSQSLSIT CTVSGFSLTT YGVHWVRQSP GKGLEWLGVM WRGGSTDFNA AFMSRLSIT KDNSKSQVFF KMNSLQADDT AIYYCARYGN YDAMDYWGQG TSVTVSSAKT TPPSVYPLAP GSAAQTNSMV T LGCLVKGY ...String: MAVLVLLLCL VTFPSCVLSQ VQLKQSGPGL VQPSQSLSIT CTVSGFSLTT YGVHWVRQSP GKGLEWLGVM WRGGSTDFNA AFMSRLSIT KDNSKSQVFF KMNSLQADDT AIYYCARYGN YDAMDYWGQG TSVTVSSAKT TPPSVYPLAP GSAAQTNSMV T LGCLVKGY FPEPVTVTWN SGSLSSGVHT FPAVLQSDLY TLSSSVTVPS SPRPSETVTC NVAHPASSTK VDKKIVPRDC |
-Macromolecule #6: CARDIOLIPIN
| Macromolecule | Name: CARDIOLIPIN / type: ligand / ID: 6 / Number of copies: 1 / Formula: CDL |
|---|---|
| Molecular weight | Theoretical: 1.464043 KDa |
| Chemical component information |  ChemComp-CDL: |
-Macromolecule #7: PHOSPHATIDYLETHANOLAMINE
| Macromolecule | Name: PHOSPHATIDYLETHANOLAMINE / type: ligand / ID: 7 / Number of copies: 1 / Formula: PTY |
|---|---|
| Molecular weight | Theoretical: 734.039 Da |
| Chemical component information |  ChemComp-PTY: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)