[English] 日本語
 Yorodumi
Yorodumi- EMDB-27753: CryoEM structure of Gq-coupled MRGPRX1 with peptide ligand BAM8-2... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of Gq-coupled MRGPRX1 with peptide ligand BAM8-22 and positive allosteric modulator ML382 | |||||||||
 Map data Map data | CryoEM structure of Gq-coupled MRGPRX1 with peptide ligand BAM8-22 and positive allosteric modulator ML382 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / SIGNALING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationPeptide ligand-binding receptors / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / Post-translational protein phosphorylation / response to chloroquine / chromaffin granule lumen / opioid peptide activity / aggressive behavior / G protein-coupled opioid receptor signaling pathway / sensory perception / G alpha (i) signalling events ...Peptide ligand-binding receptors / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / Post-translational protein phosphorylation / response to chloroquine / chromaffin granule lumen / opioid peptide activity / aggressive behavior / G protein-coupled opioid receptor signaling pathway / sensory perception / G alpha (i) signalling events / startle response / transmission of nerve impulse / neuropeptide signaling pathway / behavioral fear response / axon terminus / sensory perception of pain / acute-phase response / locomotory behavior / G protein-coupled receptor activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G protein-coupled acetylcholine receptor signaling pathway / G-protein activation / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through CDC42 / Glucagon signaling in metabolic regulation / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / ADP signalling through P2Y purinoceptor 12 / photoreceptor disc membrane / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / ADP signalling through P2Y purinoceptor 1 / cellular response to catecholamine stimulus / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / transmembrane signaling receptor activity / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / adenylate cyclase-activating dopamine receptor signaling pathway / GPER1 signaling / Inactivation, recovery and regulation of the phototransduction cascade / cellular response to prostaglandin E stimulus / G-protein beta-subunit binding / heterotrimeric G-protein complex / G alpha (12/13) signalling events / sensory perception of taste / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / retina development in camera-type eye / GTPase binding / Ca2+ pathway / fibroblast proliferation / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / chemical synaptic transmission / G alpha (q) signalling events / Ras protein signal transduction / Extra-nuclear estrogen signaling / cell surface receptor signaling pathway / cell population proliferation / defense response to bacterium / G protein-coupled receptor signaling pathway / lysosomal membrane / neuronal cell body / GTPase activity / synapse / dendrite / protein-containing complex binding / cell surface / signal transduction / extracellular exosome / extracellular region / membrane / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.71 Å | |||||||||
 Authors Authors | Liu Y / Cao C / Fay JF / Roth BL | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Chem Biol / Year: 2023 Journal: Nat Chem Biol / Year: 2023Title: Ligand recognition and allosteric modulation of the human MRGPRX1 receptor. Authors: Yongfeng Liu / Can Cao / Xi-Ping Huang / Ryan H Gumpper / Moira M Rachman / Sheng-Luen Shih / Brian E Krumm / Shicheng Zhang / Brian K Shoichet / Jonathan F Fay / Bryan L Roth /  Abstract: The human MAS-related G protein-coupled receptor X1 (MRGPRX1) is preferentially expressed in the small-diameter primary sensory neurons and involved in the mediation of nociception and pruritus. ...The human MAS-related G protein-coupled receptor X1 (MRGPRX1) is preferentially expressed in the small-diameter primary sensory neurons and involved in the mediation of nociception and pruritus. Central activation of MRGPRX1 by the endogenous opioid peptide fragment BAM8-22 and its positive allosteric modulator ML382 has been shown to effectively inhibit persistent pain, making MRGPRX1 a promising target for non-opioid pain treatment. However, the activation mechanism of MRGPRX1 is still largely unknown. Here we report three high-resolution cryogenic electron microscopy structures of MRGPRX1-Gαq in complex with BAM8-22 alone, with BAM8-22 and ML382 simultaneously as well as with a synthetic agonist compound-16. These structures reveal the agonist binding mode for MRGPRX1 and illuminate the structural requirements for positive allosteric modulation. Collectively, our findings provide a molecular understanding of the activation and allosteric modulation of the MRGPRX1 receptor, which could facilitate the structure-based design of non-opioid pain-relieving drugs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27753.map.gz emd_27753.map.gz | 77.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27753-v30.xml emd-27753-v30.xml emd-27753.xml emd-27753.xml | 27.5 KB 27.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27753_fsc.xml emd_27753_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_27753.png emd_27753.png | 73.3 KB | ||
| Masks |  emd_27753_msk_1.map emd_27753_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27753.cif.gz emd-27753.cif.gz | 7.6 KB | ||
| Others |  emd_27753_additional_1.map.gz emd_27753_additional_1.map.gz emd_27753_half_map_1.map.gz emd_27753_half_map_1.map.gz emd_27753_half_map_2.map.gz emd_27753_half_map_2.map.gz | 85.2 MB 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27753 http://ftp.pdbj.org/pub/emdb/structures/EMD-27753 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27753 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27753 | HTTPS FTP |
-Validation report
| Summary document |  emd_27753_validation.pdf.gz emd_27753_validation.pdf.gz | 816.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27753_full_validation.pdf.gz emd_27753_full_validation.pdf.gz | 815.9 KB | Display | |
| Data in XML |  emd_27753_validation.xml.gz emd_27753_validation.xml.gz | 18.7 KB | Display | |
| Data in CIF |  emd_27753_validation.cif.gz emd_27753_validation.cif.gz | 24.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27753 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27753 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27753 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27753 | HTTPS FTP |
-Related structure data
| Related structure data |  8dwgMC  8dwcC  8dwhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27753.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27753.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of Gq-coupled MRGPRX1 with peptide ligand BAM8-22 and positive allosteric modulator ML382 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : MRGPRX1-Gq BAM8-22
| Entire | Name: MRGPRX1-Gq BAM8-22 |
|---|---|
| Components |
|
-Supramolecule #1: MRGPRX1-Gq BAM8-22
| Supramolecule | Name: MRGPRX1-Gq BAM8-22 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 150 kDa/nm |
-Macromolecule #1: Proenkephalin-A
| Macromolecule | Name: Proenkephalin-A / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.974225 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VGRPEWWMDY QKRYG UniProtKB: Proenkephalin-A |
-Macromolecule #2: Gs-mini-Gq chimera
| Macromolecule | Name: Gs-mini-Gq chimera / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.084832 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSTVSAEDK AAAERSKMID KNLREDGEKA RRTLRLLLLG ADNSGKSTIV KQMRILHGGS GGSGGTSGIF ETKFQVDKVN FHMFDVGGQ RDERRKWIQC FNDVTAIIFV VDSSDYNRLQ EALNDFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK I EDYFPEFA ...String: MGSTVSAEDK AAAERSKMID KNLREDGEKA RRTLRLLLLG ADNSGKSTIV KQMRILHGGS GGSGGTSGIF ETKFQVDKVN FHMFDVGGQ RDERRKWIQC FNDVTAIIFV VDSSDYNRLQ EALNDFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK I EDYFPEFA RYTTPEDATP EPGEDPRVTR AKYFIRKEFV DISTASGDGR HICYPHFTCA VDTENARRIF NDCKDIILQM NL REYNLV |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.728152 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPGSSGSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: GPGSSGSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: Mas-related G-protein coupled receptor member X1
| Macromolecule | Name: Mas-related G-protein coupled receptor member X1 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 36.299879 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPDPTISTLD TELTPINGTE ETLCYKQTLS LTVLTCIVSL VGLTGNAVVL WLLGCRMRRN AFSIYILNLA AADFLFLSGR LIYSLLSFI SIPHTISKIL YPVMMFSYFA GLSFLSAVST ERCLSVLWPI WYRCHRPTHL SAVVCVLLWA LSLLRSILEW M LCGFLFSG ...String: GPDPTISTLD TELTPINGTE ETLCYKQTLS LTVLTCIVSL VGLTGNAVVL WLLGCRMRRN AFSIYILNLA AADFLFLSGR LIYSLLSFI SIPHTISKIL YPVMMFSYFA GLSFLSAVST ERCLSVLWPI WYRCHRPTHL SAVVCVLLWA LSLLRSILEW M LCGFLFSG ADSAWCQTSD FITVAWLIFL CVVLCGSSLV LLIRILCGSR KIPLTRLYVT ILLTVLVFLL CGLPFGIQFF LF LWIHVDR EVLFCHVHLV SIFLSALNSS ANPIIYFFVG SFRQRQNRQN LKLVLQRALQ DASEVDEGGG QLPEEILELS GSR LEQ UniProtKB: Mas-related G-protein coupled receptor member X1 |
-Macromolecule #6: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.409588 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAALEVLFQ |
-Macromolecule #7: 2-[(cyclopropanesulfonyl)amino]-N-(2-ethoxyphenyl)benzamide
| Macromolecule | Name: 2-[(cyclopropanesulfonyl)amino]-N-(2-ethoxyphenyl)benzamide type: ligand / ID: 7 / Number of copies: 1 / Formula: U39 |
|---|---|
| Molecular weight | Theoretical: 360.427 Da |
| Chemical component information | 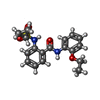 ChemComp-U39: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.2 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























