+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2750 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the CsgG-CsgE complex | |||||||||
 Map data Map data | reconstruction of the CsgG-CsgE complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Bacterial secretion / membrane protein | |||||||||
| Function / homology |  Function and homology information Function and homology information: / curli secretion complex / curli assembly / protein secretion by the type VIII secretion system / protein transmembrane transport / single-species biofilm formation / cell outer membrane / outer membrane-bounded periplasmic space / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 24.0 Å | |||||||||
 Authors Authors | Krasteva PV / Gubellini F / Remaut H / Fronzes R | |||||||||
 Citation Citation |  Journal: Nature / Year: 2014 Journal: Nature / Year: 2014Title: Structural and mechanistic insights into the bacterial amyloid secretion channel CsgG. Authors: Parveen Goyal / Petya V Krasteva / Nani Van Gerven / Francesca Gubellini / Imke Van den Broeck / Anastassia Troupiotis-Tsaïlaki / Wim Jonckheere / Gérard Péhau-Arnaudet / Jerome S Pinkner ...Authors: Parveen Goyal / Petya V Krasteva / Nani Van Gerven / Francesca Gubellini / Imke Van den Broeck / Anastassia Troupiotis-Tsaïlaki / Wim Jonckheere / Gérard Péhau-Arnaudet / Jerome S Pinkner / Matthew R Chapman / Scott J Hultgren / Stefan Howorka / Rémi Fronzes / Han Remaut /     Abstract: Curli are functional amyloid fibres that constitute the major protein component of the extracellular matrix in pellicle biofilms formed by Bacteroidetes and Proteobacteria (predominantly of the α ...Curli are functional amyloid fibres that constitute the major protein component of the extracellular matrix in pellicle biofilms formed by Bacteroidetes and Proteobacteria (predominantly of the α and γ classes). They provide a fitness advantage in pathogenic strains and induce a strong pro-inflammatory response during bacteraemia. Curli formation requires a dedicated protein secretion machinery comprising the outer membrane lipoprotein CsgG and two soluble accessory proteins, CsgE and CsgF. Here we report the X-ray structure of Escherichia coli CsgG in a non-lipidated, soluble form as well as in its native membrane-extracted conformation. CsgG forms an oligomeric transport complex composed of nine anticodon-binding-domain-like units that give rise to a 36-stranded β-barrel that traverses the bilayer and is connected to a cage-like vestibule in the periplasm. The transmembrane and periplasmic domains are separated by a 0.9-nm channel constriction composed of three stacked concentric phenylalanine, asparagine and tyrosine rings that may guide the extended polypeptide substrate through the secretion pore. The specificity factor CsgE forms a nonameric adaptor that binds and closes off the periplasmic face of the secretion channel, creating a 24,000 Å(3) pre-constriction chamber. Our structural, functional and electrophysiological analyses imply that CsgG is an ungated, non-selective protein secretion channel that is expected to employ a diffusion-based, entropy-driven transport mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2750.map.gz emd_2750.map.gz | 790.7 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2750-v30.xml emd-2750-v30.xml emd-2750.xml emd-2750.xml | 11.9 KB 11.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2750.png emd_2750.png | 70.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2750 http://ftp.pdbj.org/pub/emdb/structures/EMD-2750 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2750 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2750 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2750.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2750.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
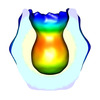
| Annotation | reconstruction of the CsgG-CsgE complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.99 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : CsgG-CsgE complex
| Entire | Name: CsgG-CsgE complex |
|---|---|
| Components |
|
-Supramolecule #1000: CsgG-CsgE complex
| Supramolecule | Name: CsgG-CsgE complex / type: sample / ID: 1000 Details: The sample was reconstituted from purified components in vitro. Oligomeric state: One nonamer of CsgG bound to one nonamer of CsgE Number unique components: 2 |
|---|---|
| Molecular weight | Theoretical: 400 KDa |
-Macromolecule #1: CsgG
| Macromolecule | Name: CsgG / type: protein_or_peptide / ID: 1 Name.synonym: Curli production assembly/transport component CsgG Details: Strep-tagged / Number of copies: 9 / Oligomeric state: nonamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 290 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Curli production assembly/transport component CsgG GO: GO: 0022610, single-species biofilm formation, protein transmembrane transport, cell outer membrane InterPro: Curli production assembly/transport component CsgG |
-Macromolecule #2: CsgE
| Macromolecule | Name: CsgE / type: protein_or_peptide / ID: 2 Name.synonym: Curli production assembly/transport component CsgE Details: Poly-histidine tagged. Used for CsgE-CsgG complex reconstitution and pull-down on Talon metal affinity resin. Number of copies: 9 / Oligomeric state: nonamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 110 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Curli production assembly/transport component CsgE GO: GO: 0022610, single-species biofilm formation, protein transmembrane transport, cell outer membrane InterPro: Curli assembly protein CsgE |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.05 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM Tris-HCl pH 8.0, 100 mM NaCl, 100 mM imidazole, 2.5% xylitol, 0.2 mM DTT |
| Grid | Details: Quantifoil R2/2 carbon coated grids (Quantifoil Micro Tools GmbH) |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 92 K / Instrument: LEICA EM GP / Method: Blot for 0.5 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Average: 95 K |
| Alignment procedure | Legacy - Astigmatism: Astigmatism was checked during imaging and during CTF correction of the micrographs. Micrographs is astigmatism >10% were discarded. |
| Date | Jun 2, 2014 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON II (4k x 4k) / Number real images: 75 / Average electron dose: 20 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 70350 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Particles were automatically selected from CTF-corrected micrographs using BOXER (EMAN2). |
|---|---|
| CTF correction | Details: Each micrograph |
| Final reconstruction | Applied symmetry - Point group: C9 (9 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 24.0 Å / Resolution method: OTHER / Software - Name: IMAGIC / Number images used: 1221 |
| Final two d classification | Number classes: 125 |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















