+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human TMEM175 in complex with 4-aminopyridine | |||||||||
 Map data Map data | hsTMEM175 in complex with 4-aminopyridine | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | potassium channel / ion channel / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationlysosomal lumen pH elevation / phagosome-lysosome fusion / : / potassium ion leak channel activity / proton channel activity / arachidonate binding / potassium channel activity / potassium ion transmembrane transport / proton transmembrane transport / neuron cellular homeostasis ...lysosomal lumen pH elevation / phagosome-lysosome fusion / : / potassium ion leak channel activity / proton channel activity / arachidonate binding / potassium channel activity / potassium ion transmembrane transport / proton transmembrane transport / neuron cellular homeostasis / lysosome / endosome / endosome membrane / lysosomal membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
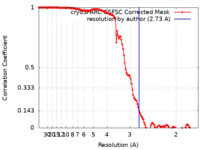
| Method | single particle reconstruction / cryo EM / Resolution: 2.73 Å | |||||||||
 Authors Authors | Oh S / Hite RK | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Mechanism of 4-aminopyridine inhibition of the lysosomal channel TMEM175. Authors: SeCheol Oh / Robyn Stix / Wenchang Zhou / José D Faraldo-Gómez / Richard K Hite /  Abstract: Transmembrane protein 175 (TMEM175) is an evolutionarily distinct lysosomal cation channel whose mutation is associated with the development of Parkinson's disease. Here, we present a cryoelectron ...Transmembrane protein 175 (TMEM175) is an evolutionarily distinct lysosomal cation channel whose mutation is associated with the development of Parkinson's disease. Here, we present a cryoelectron microscopy structure and molecular simulations of TMEM175 bound to 4-aminopyridine (4-AP), the only known small-molecule inhibitor of TMEM175 and a broad K channel inhibitor, as well as a drug approved by the Food and Drug Administration against multiple sclerosis. The structure shows that 4-AP, whose mode of action had not been previously visualized, binds near the center of the ion conduction pathway, in the open state of the channel. Molecular dynamics simulations reveal that this binding site is near the middle of the transmembrane potential gradient, providing a rationale for the voltage-dependent dissociation of 4-AP from TMEM175. Interestingly, bound 4-AP rapidly switches between three predominant binding poses, stabilized by alternate interaction patterns dictated by the twofold symmetry of the channel. Despite this highly dynamic binding mode, bound 4-AP prevents not only ion permeation but also water flow. Together, these studies provide a framework for the rational design of novel small-molecule inhibitors of TMEM175 that might reveal the role of this channel in human lysosomal physiology both in health and disease. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27436.map.gz emd_27436.map.gz | 107.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27436-v30.xml emd-27436-v30.xml emd-27436.xml emd-27436.xml | 20.4 KB 20.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27436_fsc.xml emd_27436_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_27436.png emd_27436.png | 173.1 KB | ||
| Filedesc metadata |  emd-27436.cif.gz emd-27436.cif.gz | 6.2 KB | ||
| Others |  emd_27436_additional_1.map.gz emd_27436_additional_1.map.gz emd_27436_half_map_1.map.gz emd_27436_half_map_1.map.gz emd_27436_half_map_2.map.gz emd_27436_half_map_2.map.gz | 676 MB 200.1 MB 200.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27436 http://ftp.pdbj.org/pub/emdb/structures/EMD-27436 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27436 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27436 | HTTPS FTP |
-Related structure data
| Related structure data |  8dhmMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27436.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27436.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | hsTMEM175 in complex with 4-aminopyridine | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Density modified and resampled 1.5x at
| File | emd_27436_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Density modified and resampled 1.5x at | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 1
| File | emd_27436_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_27436_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human TMEM175
| Entire | Name: Human TMEM175 |
|---|---|
| Components |
|
-Supramolecule #1: Human TMEM175
| Supramolecule | Name: Human TMEM175 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Endosomal/lysosomal potassium channel TMEM175
| Macromolecule | Name: Endosomal/lysosomal potassium channel TMEM175 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.667219 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSQPRTPEQA LDTPGDCPPG RRDEDAGEGI QCSQRMLSFS DALLSIIATV MILPVTHTEI SPEQQFDRSV QRLLATRIAV YLMTFLIVT VAWAAHTRLF QVVGKTDDTL ALLNLACMMT ITFLPYTFSL MVTFPDVPLG IFLFCVCVIA IGVVQALIVG Y AFHFPHLL ...String: MSQPRTPEQA LDTPGDCPPG RRDEDAGEGI QCSQRMLSFS DALLSIIATV MILPVTHTEI SPEQQFDRSV QRLLATRIAV YLMTFLIVT VAWAAHTRLF QVVGKTDDTL ALLNLACMMT ITFLPYTFSL MVTFPDVPLG IFLFCVCVIA IGVVQALIVG Y AFHFPHLL SPQIQRSAHR ALYRRHVLGI VLQGPALCFA AAIFSLFFVP LSYLLMVTVI LLPYVSKVTG WCRDRLLGHR EP SAHPVEV FSFDLHEPLS KERVEAFSDG VYAIVATLLI LDICEDNVPD PKDVKERFSG SLVAALSATG PRFLAYFGSF ATV GLLWFA HHSLFLHVRK ATRAMGLLNT LSLAFVGGLP LAYQQTSAFA RQPRDELERV RVSCTIIFLA SIFQLAMWTT ALLH QAETL QPSVWFGGRE HVLMFAKLAL YPCASLLAFA STCLLSRFSV GIFHLMQIAV PCAFLLLRLL VGLALATLRV LRGLA RPEH PPPAPTGQDD PQSQLLPAPC UniProtKB: Endosomal/lysosomal proton channel TMEM175 |
-Macromolecule #2: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Macromolecule #3: 4-AMINOPYRIDINE
| Macromolecule | Name: 4-AMINOPYRIDINE / type: ligand / ID: 3 / Number of copies: 1 / Formula: 4AP |
|---|---|
| Molecular weight | Theoretical: 95.122 Da |
| Chemical component information | 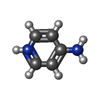 ChemComp-4AP: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 91 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 297 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 61.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)