[English] 日本語
 Yorodumi
Yorodumi- EMDB-26980: Human excitatory amino acid transporter 3 in inward facing state ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human excitatory amino acid transporter 3 in inward facing state in 100 mM L-glutamate | |||||||||
 Map data Map data | Human Excitatory amino acid transporter 3 in100 mM L-Glutamate | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
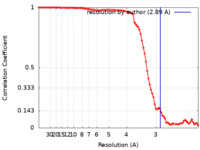
| Method | single particle reconstruction / cryo EM / Resolution: 2.89 Å | |||||||||
 Authors Authors | Qiu B / Boudker O | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Hold for publication Authors: Qiu B / Boudker O | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26980.map.gz emd_26980.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26980-v30.xml emd-26980-v30.xml emd-26980.xml emd-26980.xml | 16 KB 16 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26980_fsc.xml emd_26980_fsc.xml | 8.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_26980.png emd_26980.png | 73.1 KB | ||
| Others |  emd_26980_half_map_1.map.gz emd_26980_half_map_1.map.gz emd_26980_half_map_2.map.gz emd_26980_half_map_2.map.gz | 59.1 MB 59.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26980 http://ftp.pdbj.org/pub/emdb/structures/EMD-26980 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26980 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26980 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26980.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26980.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human Excitatory amino acid transporter 3 in100 mM L-Glutamate | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.083 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half Map 1
| File | emd_26980_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
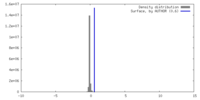
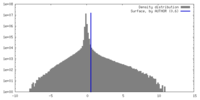
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_26980_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
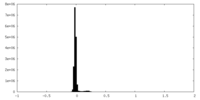
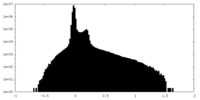
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human EAAT3 trimer in 10mM Glu
| Entire | Name: Human EAAT3 trimer in 10mM Glu |
|---|---|
| Components |
|
-Supramolecule #1: Human EAAT3 trimer in 10mM Glu
| Supramolecule | Name: Human EAAT3 trimer in 10mM Glu / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Human Excitatory amino acid transporter 3
| Macromolecule | Name: Human Excitatory amino acid transporter 3 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GPMGKPARKG CEWKRFLKNN WVLLSTVAAV VLGITTGVLV REHSNLSTLE KFYFAFPGEI LMRMLKLIIL PL IISSMIT GVAALDSNVS GKIGLRAVVY YFCTTLIAVI LGIVLVVSIK PGVTQKVGEI ARTGSTPEVS TVD AMLDLI RNMFPENLVQ ACFQQYKTKR ...String: GPMGKPARKG CEWKRFLKNN WVLLSTVAAV VLGITTGVLV REHSNLSTLE KFYFAFPGEI LMRMLKLIIL PL IISSMIT GVAALDSNVS GKIGLRAVVY YFCTTLIAVI LGIVLVVSIK PGVTQKVGEI ARTGSTPEVS TVD AMLDLI RNMFPENLVQ ACFQQYKTKR EEVKPPSDPE MTMTEESFTA VMTTAISKTK TKEYKIVGMY SDGI NVLGL IVFCLVFGLV IGKMGEKGQI LVDFFNALSD ATMKIVQIIM CYMPLGILFL IAGKIIEVED WEIFR KLGL YMATVLTGLA IHSIVILPLI YFIVVRKNPF RFAMGMAQAL LTALMISSSS ATLPVTFRCA EENNQV DKR ITRFVLPVGA TINMDGTALY EAVAAVFIAQ LNDLDLGIGQ IITISITATS ASIGAAGVPQ AGLVTMV IV LSAVGLPAED VTLIIAVDWL LDRFRTMVNV LGDAFGTGIV EKLSKKELEQ MDVSSEVNIV NPFALEST I LDNEDSDTKK SYVNGGFAVD KSDTISFTQT SQF |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Tris is used for adjusting pH | ||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot 3s. | ||||||||||||||||||
| Details | This sample was mono disperse. The glutamate concentration is 100mM, |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 51.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 2.7 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)