+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the SAGA core module | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Vasyliuk D / Yip CK | |||||||||
| Funding support |  Canada, 1 items Canada, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2022 Journal: Sci Rep / Year: 2022Title: Conformational landscape of the yeast SAGA complex as revealed by cryo-EM. Authors: Diana Vasyliuk / Joeseph Felt / Ellen D Zhong / Bonnie Berger / Joseph H Davis / Calvin K Yip /   Abstract: Spt-Ada-Gcn5-Acetyltransferase (SAGA) is a conserved multi-subunit complex that activates RNA polymerase II-mediated transcription by acetylating and deubiquitinating nucleosomal histones and by ...Spt-Ada-Gcn5-Acetyltransferase (SAGA) is a conserved multi-subunit complex that activates RNA polymerase II-mediated transcription by acetylating and deubiquitinating nucleosomal histones and by recruiting TATA box binding protein (TBP) to DNA. The prototypical yeast Saccharomyces cerevisiae SAGA contains 19 subunits that are organized into Tra1, core, histone acetyltransferase, and deubiquitination modules. Recent cryo-electron microscopy studies have generated high-resolution structural information on the Tra1 and core modules of yeast SAGA. However, the two catalytical modules were poorly resolved due to conformational flexibility of the full assembly. Furthermore, the high sample requirement created a formidable barrier to further structural investigations of SAGA. Here, we report a workflow for isolating/stabilizing yeast SAGA and preparing cryo-EM specimens at low protein concentration using a graphene oxide support layer. With this procedure, we were able to determine a cryo-EM reconstruction of yeast SAGA at 3.1 Å resolution and examine its conformational landscape with the neural network-based algorithm cryoDRGN. Our analysis revealed that SAGA adopts a range of conformations with its HAT module and central core in different orientations relative to Tra1. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26804.map.gz emd_26804.map.gz | 303.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26804-v30.xml emd-26804-v30.xml emd-26804.xml emd-26804.xml | 14.3 KB 14.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26804_fsc.xml emd_26804_fsc.xml | 14.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_26804.png emd_26804.png | 77.2 KB | ||
| Others |  emd_26804_half_map_1.map.gz emd_26804_half_map_1.map.gz emd_26804_half_map_2.map.gz emd_26804_half_map_2.map.gz | 321.4 MB 321.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26804 http://ftp.pdbj.org/pub/emdb/structures/EMD-26804 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26804 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26804 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26804.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26804.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.332 Å | ||||||||||||||||||||||||||||||||||||
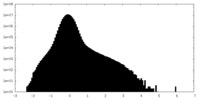
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_26804_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
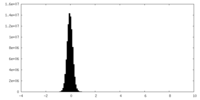
| Density Histograms |
-Half map: #2
| File | emd_26804_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
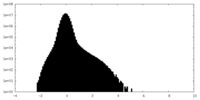
| Density Histograms |
- Sample components
Sample components
-Entire : SAGA
| Entire | Name: SAGA |
|---|---|
| Components |
|
-Supramolecule #1: SAGA
| Supramolecule | Name: SAGA / type: complex / Chimera: Yes / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: EMS Lacey Carbon / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 2s blot time, -5 blot force.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Details | PNCC Krios 4 |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 17955 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 64000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)