[English] 日本語
 Yorodumi
Yorodumi- EMDB-26644: SARS-CoV-2 Omicron-BA.2 1-RBD-up Spike Protein Trimer without the... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 Omicron-BA.2 1-RBD-up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2) | |||||||||
 Map data Map data | Sharp map from homogeneous C1 refinement in cryosparc | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Omicron Spike protein / SARS-CoV-2 / variant of concern / 1-up RBD / VIRAL PROTEIN / Omicron-BA.2 / BA.2 | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.93 Å | |||||||||
 Authors Authors | Stalls V / Acharya P | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2022 Journal: Cell Rep / Year: 2022Title: Cryo-EM structures of SARS-CoV-2 Omicron BA.2 spike. Authors: Victoria Stalls / Jared Lindenberger / Sophie M-C Gobeil / Rory Henderson / Rob Parks / Maggie Barr / Margaret Deyton / Mitchell Martin / Katarzyna Janowska / Xiao Huang / Aaron May / Micah ...Authors: Victoria Stalls / Jared Lindenberger / Sophie M-C Gobeil / Rory Henderson / Rob Parks / Maggie Barr / Margaret Deyton / Mitchell Martin / Katarzyna Janowska / Xiao Huang / Aaron May / Micah Speakman / Esther Beaudoin / Bryan Kraft / Xiaozhi Lu / Robert J Edwards / Amanda Eaton / David C Montefiori / Wilton B Williams / Kevin O Saunders / Kevin Wiehe / Barton F Haynes / Priyamvada Acharya /  Abstract: The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron BA.2 sub-lineage has gained in proportion relative to BA.1. Because spike (S) protein variations may underlie differences in ...The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron BA.2 sub-lineage has gained in proportion relative to BA.1. Because spike (S) protein variations may underlie differences in their pathobiology, here we determine cryoelectron microscopy (cryo-EM) structures of the BA.2 S ectodomain and compare these with previously determined BA.1 S structures. BA.2 receptor-binding domain (RBD) mutations induce remodeling of the RBD structure, resulting in tighter packing and improved thermostability. Interprotomer RBD interactions are enhanced in the closed (or 3-RBD-down) BA.2 S, while the fusion peptide is less accessible to antibodies than in BA.1. Binding and pseudovirus neutralization assays reveal extensive immune evasion while defining epitopes of two outer RBD face-binding antibodies, DH1044 and DH1193, that neutralize both BA.1 and BA.2. Taken together, our results indicate that stabilization of the closed state through interprotomer RBD-RBD packing is a hallmark of the Omicron variant and show differences in key functional regions in the BA.1 and BA.2 S proteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26644.map.gz emd_26644.map.gz | 117.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26644-v30.xml emd-26644-v30.xml emd-26644.xml emd-26644.xml | 16.9 KB 16.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_26644.png emd_26644.png | 46.8 KB | ||
| Filedesc metadata |  emd-26644.cif.gz emd-26644.cif.gz | 5.8 KB | ||
| Others |  emd_26644_half_map_1.map.gz emd_26644_half_map_1.map.gz emd_26644_half_map_2.map.gz emd_26644_half_map_2.map.gz | 115.7 MB 115.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26644 http://ftp.pdbj.org/pub/emdb/structures/EMD-26644 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26644 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26644 | HTTPS FTP |
-Validation report
| Summary document |  emd_26644_validation.pdf.gz emd_26644_validation.pdf.gz | 786.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_26644_full_validation.pdf.gz emd_26644_full_validation.pdf.gz | 786.2 KB | Display | |
| Data in XML |  emd_26644_validation.xml.gz emd_26644_validation.xml.gz | 14.1 KB | Display | |
| Data in CIF |  emd_26644_validation.cif.gz emd_26644_validation.cif.gz | 16.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26644 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26644 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26644 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26644 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26644.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26644.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharp map from homogeneous C1 refinement in cryosparc | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
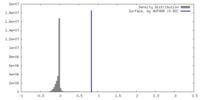
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half A map from homogeneous C1 refinement in cryosparc
| File | emd_26644_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half A map from homogeneous C1 refinement in cryosparc | ||||||||||||
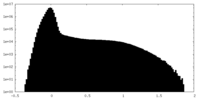
| Projections & Slices |
| ||||||||||||
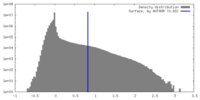
| Density Histograms |
-Half map: Half B map from homogeneous C1 refinement in cryosparc
| File | emd_26644_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half B map from homogeneous C1 refinement in cryosparc | ||||||||||||
| Projections & Slices |
| ||||||||||||
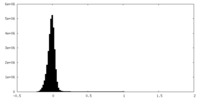
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 S-GSAS-Omicron-BA.2 Spike Ectodomain
| Entire | Name: SARS-CoV-2 S-GSAS-Omicron-BA.2 Spike Ectodomain |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 S-GSAS-Omicron-BA.2 Spike Ectodomain
| Supramolecule | Name: SARS-CoV-2 S-GSAS-Omicron-BA.2 Spike Ectodomain / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 427 KDa |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: other / ID: 1 / Classification: other |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPV LPFNDGVYFA STEKSNIIRG WIFGTTLDSK TQSLLIVNNA TNVVIKVCEF QFCNDPFLDV YYHKNNKSWM ESEFRVYSSA NNCTFEYVSQ ...String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPV LPFNDGVYFA STEKSNIIRG WIFGTTLDSK TQSLLIVNNA TNVVIKVCEF QFCNDPFLDV YYHKNNKSWM ESEFRVYSSA NNCTFEYVSQ PFLMDLEGKQ GNFKNLREFV FKNIDGYFKI YSKHTPINLG RDLPQGFSAL EPLVDLPIGI NITRFQTLLA LHRSYLTPGD SSSGWTAGAA AYYVGYLQPR TFLLKYNENG TITDAVDCAL DPLSETKCTL KSFTVEKGIY QTSNFRVQPT ESIVRFPNIT NLCPFDEVFN ATRFASVYAW NRKRISNCVA DYSVLYNFAP FFAFKCYGVS PTKLNDLCFT NVYADSFVIR GNEVSQIAPG QTGNIADYNY KLPDDFTGCV IAWNSNKLDS KVGGNYNYLY RLFRKSNLKP FERDISTEIY QAGNKPCNGV AGFNCYFPLR SYGFRPTYGV GHQPYRVVVL SFELLHAPAT VCGPKKSTNL VKNKCVNFNF NGLTGTGVLT ESNKKFLPFQ QFGRDIADTT DAVRDPQTLE ILDITPCSFG GVSVITPGTN TSNQVAVLYQ GVNCTEVPVA IHADQLTPTW RVYSTGSNVF QTRAGCLIGA EYVNNSYECD IPIGAGICAS YQTQTKSHGS ASSVASQSII AYTMSLGAEN SVAYSNNSIA IPTNFTISVT TEILPVSMTK TSVDCTMYIC GDSTECSNLL LQYGSFCTQL KRALTGIAVE QDKNTQEVFA QVKQIYKTPP IKYFGGFNFS QILPDPSKPS KRSFIEDLLF NKVTLADAGF IKQYGDCLGD IAARDLICAQ KFNGLTVLPP LLTDEMIAQY TSALLAGTIT SGWTFGAGAA LQIPFAMQMA YRFNGIGVTQ NVLYENQKLI ANQFNSAIGK IQDSLSSTAS ALGKLQDVVN HNAQALNTLV KQLSSKFGAI SSVLNDILSR LDKVEAEVQI DRLITGRLQS LQTYVTQQLI RAAEIRASAN LAATKMSECV LGQSKRVDFC GKGYHLMSFP QSAPHGVVFL HVTYVPAQEK NFTTAPAICH DGKAHFPREG VFVSNGTHWF VTQRNFYEPQ IITTDNTFVS GNCDVVIGIV NNTVYDPLQP ELDSFKEELD KYFKNHTSPD VDLGDISGIN ASVVNIQKEI DRLNEVAKNL NESLIDLQEL GKYEQGSGYI PEAPRDGQAY VRKDGEWVLL STFLGRSLEV LFQGPGHHHH HHHHSAWSHP QFEKGGGSGG GGSGGSAWSH PQFEK |
| Recombinant expression | Organism: Mammalia (mammals) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 11742 / Average electron dose: 54.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.6 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)