+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Pol II-DSIF-SPT6-PAF1c-TFIIS complex with rewrapped nucleosome | |||||||||
 マップデータ マップデータ | Map E, 54 complex, composite | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Chromatin / nucleosome / transcription / intermediate | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報blastocyst growth / Ski complex / RNA polymerase II C-terminal domain phosphoserine binding / mRNA decay by 3' to 5' exoribonuclease / positive regulation of mRNA 3'-end processing / Cdc73/Paf1 complex / inner cell mass cell differentiation / regulation of isotype switching / negative regulation of DNA-templated transcription, elongation / nuclear-transcribed mRNA catabolic process, 3'-5' exonucleolytic nonsense-mediated decay ...blastocyst growth / Ski complex / RNA polymerase II C-terminal domain phosphoserine binding / mRNA decay by 3' to 5' exoribonuclease / positive regulation of mRNA 3'-end processing / Cdc73/Paf1 complex / inner cell mass cell differentiation / regulation of isotype switching / negative regulation of DNA-templated transcription, elongation / nuclear-transcribed mRNA catabolic process, 3'-5' exonucleolytic nonsense-mediated decay / regulation of muscle cell differentiation / endodermal cell fate commitment / negative regulation of myeloid cell differentiation / positive regulation of cell cycle G1/S phase transition / DSIF complex / trophectodermal cell differentiation / regulation of transcription elongation by RNA polymerase II / blastocyst hatching / nucleosome organization / blastocyst formation / nuclear lumen / mRNA 3'-end processing / positive regulation of DNA-templated transcription, elongation / Abortive elongation of HIV-1 transcript in the absence of Tat / stem cell population maintenance / transcription factor TFIID complex / interleukin-6-mediated signaling pathway / negative regulation of G1/S transition of mitotic cell cycle / transcription elongation-coupled chromatin remodeling / RNA Pol II CTD phosphorylation and interaction with CE during HIV infection / RNA Pol II CTD phosphorylation and interaction with CE / negative regulation of gene expression, epigenetic / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping / RNA polymerase II complex binding / Pausing and recovery of Tat-mediated HIV elongation / Tat-mediated HIV elongation arrest and recovery / positive regulation of macroautophagy / RNA polymerase II transcribes snRNA genes / HIV elongation arrest and recovery / Pausing and recovery of HIV elongation / positive regulation of Wnt signaling pathway / protein localization to nucleus / cell surface receptor signaling pathway via JAK-STAT / mRNA transport / Tat-mediated elongation of the HIV-1 transcript / Formation of HIV-1 elongation complex containing HIV-1 Tat / RNA polymerase I complex / Formation of HIV elongation complex in the absence of HIV Tat / RNA polymerase III complex / negative regulation of fibroblast proliferation / RNA polymerase II, core complex / nucleosome binding / tRNA transcription by RNA polymerase III / RNA Polymerase II Transcription Elongation / transcription by RNA polymerase I / Formation of RNA Pol II elongation complex / RNA Polymerase II Pre-transcription Events / DNA-directed RNA polymerase complex / rescue of stalled ribosome / SH2 domain binding / RNA splicing / transcription elongation factor complex / erythrocyte differentiation / Hedgehog 'on' state / TP53 Regulates Transcription of DNA Repair Genes / transcription elongation by RNA polymerase II / DNA-templated transcription initiation / regulation of cell growth / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / positive regulation of transcription elongation by RNA polymerase II / Formation of TC-NER Pre-Incision Complex / Formation of the beta-catenin:TCF transactivating complex / euchromatin / protein destabilization / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / ribonucleoside binding / Wnt signaling pathway / fibrillar center / mRNA processing / DNA-directed RNA polymerase / negative regulation of epithelial cell proliferation / DNA-directed RNA polymerase activity / structural constituent of chromatin / E3 ubiquitin ligases ubiquitinate target proteins / nucleosome / heterochromatin formation / nucleosome assembly / single-stranded DNA binding / chromatin organization / cellular response to lipopolysaccharide / histone binding / nucleic acid binding / transcription by RNA polymerase II / chromosome, telomeric region / protein dimerization activity / nuclear speck / protein heterodimerization activity 類似検索 - 分子機能 | |||||||||
| 生物種 |   Homo sapiens (ヒト) / Homo sapiens (ヒト) / | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.0 Å | |||||||||
 データ登録者 データ登録者 | Filipovski M / Vos SM / Farnung L | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Science / 年: 2022 ジャーナル: Science / 年: 2022タイトル: Structural basis of nucleosome retention during transcription elongation. 著者: Martin Filipovski / Jelly H M Soffers / Seychelle M Vos / Lucas Farnung /  要旨: In eukaryotes, RNA polymerase (Pol) II transcribes chromatin and must move past nucleosomes, often resulting in nucleosome displacement. How Pol II unwraps the DNA from nucleosomes to allow ...In eukaryotes, RNA polymerase (Pol) II transcribes chromatin and must move past nucleosomes, often resulting in nucleosome displacement. How Pol II unwraps the DNA from nucleosomes to allow transcription and how DNA rewraps to retain nucleosomes has been unclear. Here, we report the 3.0-angstrom cryo-electron microscopy structure of a mammalian Pol II-DSIF-SPT6-PAF1c-TFIIS-nucleosome complex stalled 54 base pairs within the nucleosome. The structure provides a mechanistic basis for nucleosome retention during transcription elongation where upstream DNA emerging from the Pol II cleft has rewrapped the proximal side of the nucleosome. The structure uncovers a direct role for Pol II and transcription elongation factors in nucleosome retention and explains how nucleosomes are retained to prevent the disruption of chromatin structure across actively transcribed genes. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_26620.map.gz emd_26620.map.gz | 204.4 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-26620-v30.xml emd-26620-v30.xml emd-26620.xml emd-26620.xml | 84.3 KB 84.3 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_26620.png emd_26620.png | 158.2 KB | ||
| マスクデータ |  emd_26620_msk_1.map emd_26620_msk_1.map emd_26620_msk_2.map emd_26620_msk_2.map emd_26620_msk_3.map emd_26620_msk_3.map | 347.6 MB 347.6 MB 347.6 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-26620.cif.gz emd-26620.cif.gz | 17 KB | ||
| その他 |  emd_26620_additional_1.map.gz emd_26620_additional_1.map.gz emd_26620_additional_10.map.gz emd_26620_additional_10.map.gz emd_26620_additional_11.map.gz emd_26620_additional_11.map.gz emd_26620_additional_12.map.gz emd_26620_additional_12.map.gz emd_26620_additional_13.map.gz emd_26620_additional_13.map.gz emd_26620_additional_14.map.gz emd_26620_additional_14.map.gz emd_26620_additional_15.map.gz emd_26620_additional_15.map.gz emd_26620_additional_16.map.gz emd_26620_additional_16.map.gz emd_26620_additional_2.map.gz emd_26620_additional_2.map.gz emd_26620_additional_3.map.gz emd_26620_additional_3.map.gz emd_26620_additional_4.map.gz emd_26620_additional_4.map.gz emd_26620_additional_5.map.gz emd_26620_additional_5.map.gz emd_26620_additional_6.map.gz emd_26620_additional_6.map.gz emd_26620_additional_7.map.gz emd_26620_additional_7.map.gz emd_26620_additional_8.map.gz emd_26620_additional_8.map.gz emd_26620_additional_9.map.gz emd_26620_additional_9.map.gz | 172.8 MB 327.4 MB 328.6 MB 169 MB 328.3 MB 323 MB 322.3 MB 173.7 MB 173.8 MB 322.3 MB 323 MB 328.2 MB 322.7 MB 322.7 MB 322.2 MB 322.2 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26620 http://ftp.pdbj.org/pub/emdb/structures/EMD-26620 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26620 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26620 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_26620_validation.pdf.gz emd_26620_validation.pdf.gz | 492.1 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_26620_full_validation.pdf.gz emd_26620_full_validation.pdf.gz | 491.7 KB | 表示 | |
| XML形式データ |  emd_26620_validation.xml.gz emd_26620_validation.xml.gz | 7.6 KB | 表示 | |
| CIF形式データ |  emd_26620_validation.cif.gz emd_26620_validation.cif.gz | 8.8 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26620 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26620 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26620 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26620 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7uncMC  7undC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_26620.map.gz / 形式: CCP4 / 大きさ: 347.6 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_26620.map.gz / 形式: CCP4 / 大きさ: 347.6 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Map E, 54 complex, composite | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
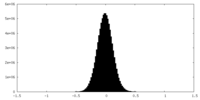
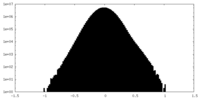
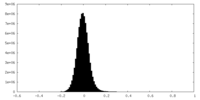
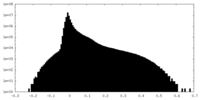
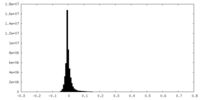
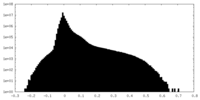
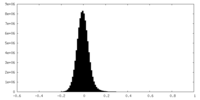
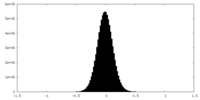
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
+マスク #1
+マスク #2
+マスク #3
+追加マップ: Map C, 54 complex, nucleosome local refinement, consensus
+追加マップ: Map D, 54 complex, CTR9/WDR61 local refinement, sharp
+追加マップ: Map A, 54 complex, overall, sharp
+追加マップ: Map D, 54 complex, CTR9/WDR61 local refinement, consensus
+追加マップ: Map B, 54 complex, Pol II (active site) local refinement, sharp
+追加マップ: Map B, 54 complex, Pol II (active site) local refinement, half A
+追加マップ: Map A, 54 complex, overall, half B
+追加マップ: Map A, 54 complex, overall, consensus
+追加マップ: Map B, 54 complex, Pol II local refinement, consensus
+追加マップ: Map A, 54 complex, overall, half A
+追加マップ: Map B, 54 complex, Pol II local refinement, half B
+追加マップ: Map C, 54 complex, nucleosome local refinement, sharp
+追加マップ: Map D, 54 complex, CTR9/WDR61 local refinement, half A
+追加マップ: Map D, 54 complex, CTR9/WDR61 local refinement, half B
+追加マップ: Map C, 54 complex, nucleosome local refinement, half B
+追加マップ: Map C, 54 complex, nucleosome local refinement, half A
- 試料の構成要素
試料の構成要素
+全体 : Pol II-DSIF-SPT6-PAF1c-TFIIS complex with rewrapped nucleosome
+超分子 #1: Pol II-DSIF-SPT6-PAF1c-TFIIS complex with rewrapped nucleosome
+分子 #1: DNA-directed RNA polymerase subunit
+分子 #2: DNA-directed RNA polymerase subunit beta
+分子 #3: DNA-directed RNA polymerase II subunit RPB3
+分子 #4: RPOL4c domain-containing protein
+分子 #5: DNA-directed RNA polymerase II subunit E
+分子 #6: DNA-directed RNA polymerases I, II, and III subunit RPABC2
+分子 #7: DNA-directed RNA polymerase II subunit RPB7
+分子 #8: RPB8
+分子 #9: RPB9
+分子 #10: RPB10
+分子 #11: RPB11
+分子 #12: RNA polymerase II subunit K
+分子 #13: Transcription elongation factor SPT6
+分子 #15: Transcription elongation factor A protein 1
+分子 #17: RNA polymerase-associated protein CTR9 homolog
+分子 #18: RNA polymerase-associated protein RTF1 homolog
+分子 #20: RNA polymerase-associated protein LEO1
+分子 #21: RNA polymerase II-associated factor 1 homolog
+分子 #22: WDR61
+分子 #23: Parafibromin
+分子 #24: Transcription elongation factor SPT5
+分子 #25: Histone H3.2
+分子 #26: Histone H4
+分子 #27: Histone H2A
+分子 #28: Histone H2B 1.1
+分子 #14: Non-template DNA
+分子 #19: Template DNA
+分子 #16: RNA
+分子 #29: ZINC ION
+分子 #30: MAGNESIUM ION
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 構成要素:
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| グリッド | モデル: Quantifoil R2/1 / 材質: COPPER / メッシュ: 200 / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 時間: 30 sec. / 詳細: 15 mA with 10 s hold time | ||||||||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 特殊光学系 | 位相板: VOLTA PHASE PLATE / エネルギーフィルター - スリット幅: 20 eV |
| 撮影 | フィルム・検出器のモデル: GATAN K3 BIOQUANTUM (6k x 4k) 平均電子線量: 52.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: SPOT SCAN / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.0 µm / 最小 デフォーカス(公称値): 0.5 µm / 倍率(公称値): 105000 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
- 画像解析
画像解析
| 初期モデル | モデルのタイプ: OTHER / 詳細: Ab initio |
|---|---|
| 最終 再構成 | 使用したクラス数: 1 / 解像度のタイプ: BY AUTHOR / 解像度: 3.0 Å / 解像度の算出法: FSC 0.143 CUT-OFF / 使用した粒子像数: 105420 |
| 初期 角度割当 | タイプ: MAXIMUM LIKELIHOOD |
| 最終 角度割当 | タイプ: MAXIMUM LIKELIHOOD |
 ムービー
ムービー コントローラー
コントローラー

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
















































































































































 Trichoplusia ni (イラクサキンウワバ)
Trichoplusia ni (イラクサキンウワバ)


