+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Glutamine Synthetase Type III from Ostreococcus tauri | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | hexamer / glutamine / LIGASE | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |  Ostreococcus tauri (plant) Ostreococcus tauri (plant) | |||||||||
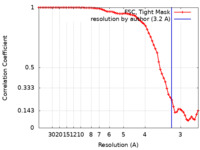
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Novikova IV / Powell SM / Evans JE | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Eukaryotic Glutamine Synthetase Type III Assembles as a Hexamer and Lacks Ring-Ring Quaternary Contacts. Authors: Novikova IV / Powell SM / Smallwood CR / Purvine SO / Zhou M / Evans JE | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26377.map.gz emd_26377.map.gz | 15.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26377-v30.xml emd-26377-v30.xml emd-26377.xml emd-26377.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26377_fsc.xml emd_26377_fsc.xml | 7.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_26377.png emd_26377.png | 116.1 KB | ||
| Filedesc metadata |  emd-26377.cif.gz emd-26377.cif.gz | 5.8 KB | ||
| Others |  emd_26377_half_map_1.map.gz emd_26377_half_map_1.map.gz emd_26377_half_map_2.map.gz emd_26377_half_map_2.map.gz | 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26377 http://ftp.pdbj.org/pub/emdb/structures/EMD-26377 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26377 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26377 | HTTPS FTP |
-Related structure data
| Related structure data |  7u6oMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26377.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26377.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3018 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_26377_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
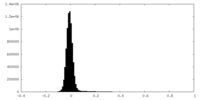
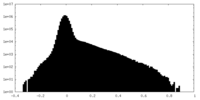
| Density Histograms |
-Half map: #2
| File | emd_26377_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
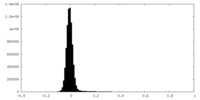
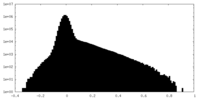
| Density Histograms |
- Sample components
Sample components
-Entire : A hexamer complex of eukaryotic glutamine synthetase type III
| Entire | Name: A hexamer complex of eukaryotic glutamine synthetase type III |
|---|---|
| Components |
|
-Supramolecule #1: A hexamer complex of eukaryotic glutamine synthetase type III
| Supramolecule | Name: A hexamer complex of eukaryotic glutamine synthetase type III type: cell / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Ostreococcus tauri (plant) Ostreococcus tauri (plant) |
-Macromolecule #1: Glutamine synthetase
| Macromolecule | Name: Glutamine synthetase / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO / EC number: glutamine synthetase |
|---|---|
| Source (natural) | Organism:  Ostreococcus tauri (plant) Ostreococcus tauri (plant) |
| Molecular weight | Theoretical: 80.09807 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDYKDHDGDY KDHDIDYKDD DDKLALSMSS GPATTEGFGS ACFKGAVADK YLSKYGESST LLANGKWTKD MAKADIVAKA VLDWAVENG ASVYCHWFQP MGSSGVRHGN SGQVHQSMFN FAEDGTPYYS FTGEQLLQGE TDGSSFPNGG MRATHTAGGY L SIDPYSPI ...String: MDYKDHDGDY KDHDIDYKDD DDKLALSMSS GPATTEGFGS ACFKGAVADK YLSKYGESST LLANGKWTKD MAKADIVAKA VLDWAVENG ASVYCHWFQP MGSSGVRHGN SGQVHQSMFN FAEDGTPYYS FTGEQLLQGE TDGSSFPNGG MRATHTAGGY L SIDPYSPI FLREDTVFIP AAFVSYNGDA LDEKTPLHRA TDALDKQTKR MLKAMKYDVG SASVYANIGL EQEIFLTPRH AF YRRPDLQ FTGRTITGKF PARGQEMSDH YMAPISRATG AFECMRQIQQ ECFKMGIPLK TRHREVAPNQ YEFAPMFGNA ISQ VDQNLM IMQVIEEVAS EHGLAALLQE KPFAGVNGSG KHNNWSIGTS DGLNLMNPKQ VNAKTGNPEI FPLVMAAMVS AVDK HGDLM RAAIASPGND FRLGAMEAPP AVMSTYLGPS LTEFLNTVKN GSLGEYAPKK KPLEFGSDTL PSIEVPAEDR NRTSP FPYG GNRFEFRAAG SSQNVSLVNT VLNTIAAEAF KIVADRLEAG EKPLAIAQDL LKTHDKCIFN GNGYDPAWPD EAVKRG IWR IDAGCDAINE LDSAKNVTLF EGMGIFTARE IQARKSVLLG HYVGSVEMEA LTMIDMINQH VIPSVKKADL GNPSKLV DA VKTIKGAVAQ IHGTEDEHKA ATLARTLRLT TMVAIREIID EFESRCPPED WTLATYSELL FFDTYPESEY GCGGGGSH H HHHHHHHH UniProtKB: Glutamine synthetase type III N terminal-domain-containing protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.2 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 BASE (4k x 4k) / Average electron dose: 100.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)