[English] 日本語
 Yorodumi
Yorodumi- EMDB-2600: Cryo-EM study of the chromatin fiber reveals a double helix twist... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2600 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM study of the chromatin fiber reveals a double helix twisted by tetra-nucleosomal units | |||||||||
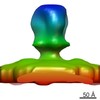
 Map data Map data | Reconstruction of 12x177 bp chromatin | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | In vitro reconstituted 12x177 bp chromatin | |||||||||
| Biological species |  Homo sapiens (human) / unidentified (others) Homo sapiens (human) / unidentified (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 11.0 Å | |||||||||
 Authors Authors | Song F / Chen P / Sun D / Wang M / Dong L / Liang D / Xu RM / Zhu P / Li G | |||||||||
 Citation Citation |  Journal: Science / Year: 2014 Journal: Science / Year: 2014Title: Cryo-EM study of the chromatin fiber reveals a double helix twisted by tetranucleosomal units. Authors: Feng Song / Ping Chen / Dapeng Sun / Mingzhu Wang / Liping Dong / Dan Liang / Rui-Ming Xu / Ping Zhu / Guohong Li /  Abstract: The hierarchical packaging of eukaryotic chromatin plays a central role in transcriptional regulation and other DNA-related biological processes. Here, we report the 11-angstrom-resolution cryogenic ...The hierarchical packaging of eukaryotic chromatin plays a central role in transcriptional regulation and other DNA-related biological processes. Here, we report the 11-angstrom-resolution cryogenic electron microscopy (cryo-EM) structures of 30-nanometer chromatin fibers reconstituted in the presence of linker histone H1 and with different nucleosome repeat lengths. The structures show a histone H1-dependent left-handed twist of the repeating tetranucleosomal structural units, within which the four nucleosomes zigzag back and forth with a straight linker DNA. The asymmetric binding and the location of histone H1 in chromatin play a role in the formation of the 30-nanometer fiber. Our results provide mechanistic insights into how nucleosomes compact into higher-order chromatin fibers. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2600.map.gz emd_2600.map.gz | 10.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2600-v30.xml emd-2600-v30.xml emd-2600.xml emd-2600.xml | 10.5 KB 10.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2600.png emd_2600.png emd_2600_1.png emd_2600_1.png | 215 KB 205.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2600 http://ftp.pdbj.org/pub/emdb/structures/EMD-2600 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2600 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2600 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2600.map.gz / Format: CCP4 / Size: 162.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2600.map.gz / Format: CCP4 / Size: 162.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of 12x177 bp chromatin | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.539 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : In vitro reconstituted 12x177 bp chromatin
| Entire | Name: In vitro reconstituted 12x177 bp chromatin |
|---|---|
| Components |
|
-Supramolecule #1000: In vitro reconstituted 12x177 bp chromatin
| Supramolecule | Name: In vitro reconstituted 12x177 bp chromatin / type: sample / ID: 1000 / Oligomeric state: Dodecamer / Number unique components: 3 |
|---|
-Macromolecule #1: Core histone
| Macromolecule | Name: Core histone / type: protein_or_peptide / ID: 1 Details: Octameric nucleosome core histone contains 2 copies histone H2A, H2B, H3 and H4. 12 octamer units constitutes the dodecamer. Number of copies: 2 / Oligomeric state: Octamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism: |
| Recombinant expression | Organism:  |
-Macromolecule #2: Histone H1.4
| Macromolecule | Name: Histone H1.4 / type: protein_or_peptide / ID: 2 / Name.synonym: HISTIHIE Details: Each octamer contains 1 copy linker histone, histone H1.4 Number of copies: 12 / Oligomeric state: Monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Location in cell: Nucleus Homo sapiens (human) / synonym: Human / Location in cell: Nucleus |
| Recombinant expression | Organism:  |
-Macromolecule #3: 601 DNA
| Macromolecule | Name: 601 DNA / type: dna / ID: 3 / Details: 12 tandem repeats; full length 2124 bp / Classification: DNA / Structure: DOUBLE HELIX / Synthetic?: No |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Sequence | String: GAGCATCCGG ATCCCCTGGA GAATCCCGGT GCCGAGGCCG CTCAATTGGT CGTAGACAGC TCTAGCACCG CTTAAACGCA CGTACGCGCT GTCCCCCGCG TTTTAACCGC CAAGGGGATT ACTCCCTAGT CTCCAGGCAC GTGTCACATA TATACATCCT GTTCCAGTGC CGGACCC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Details: 10 mM HEPES, pH 8.0, 0.1 mM EDTA |
|---|---|
| Grid | Details: 300 mesh R2.1 Quantifoil holey grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV Method: Sample absorbed for 1 to 1.5 min, blotted for 4 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 155,000 times magnification |
| Details | Parallel beam illumination |
| Date | Mar 1, 2012 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 4424 / Average electron dose: 18 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.95 µm / Nominal defocus min: 1.6 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: CTF correction of each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 11.0 Å / Resolution method: OTHER / Software - Name: EMAN2 / Number images used: 21000 |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















