[English] 日本語
 Yorodumi
Yorodumi- EMDB-25513: Cryo-EM structure of the SARS-CoV-2 D614G,N501Y mutant spike prot... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25513 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the SARS-CoV-2 D614G,N501Y mutant spike protein ectodomain bound to human ACE2 ectodomain (global refinement) | |||||||||
 Map data Map data | Global refinement map of SARS-CoV-2 D614G,N501Y mutant spike protein ectodomain bound to human ACE2 ectodomain | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / glycoprotein / fusion protein / viral protein / ACE2 / Viral Protein-Hydrolase complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / maternal process involved in female pregnancy / regulation of cardiac conduction ...positive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / maternal process involved in female pregnancy / regulation of cardiac conduction / peptidyl-dipeptidase activity / transporter activator activity / regulation of vasoconstriction / Metabolism of Angiotensinogen to Angiotensins / carboxypeptidase activity / angiotensin maturation / viral life cycle / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / metallocarboxypeptidase activity / positive regulation of cardiac muscle contraction / regulation of cytokine production / blood vessel diameter maintenance / negative regulation of smooth muscle cell proliferation / brush border membrane / negative regulation of ERK1 and ERK2 cascade / positive regulation of reactive oxygen species metabolic process / metallopeptidase activity / endocytic vesicle membrane / regulation of cell population proliferation / virus receptor activity / regulation of inflammatory response / symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / endopeptidase activity / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / Potential therapeutics for SARS / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / apical plasma membrane / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / cilium / membrane raft / endocytosis involved in viral entry into host cell / endoplasmic reticulum lumen / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / cell surface / negative regulation of transcription by RNA polymerase II / : / extracellular exosome / extracellular region / zinc ion binding / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
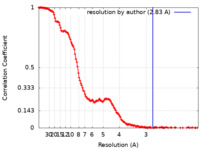
| Method | single particle reconstruction / cryo EM / Resolution: 2.83 Å | |||||||||
 Authors Authors | Zhu X / Mannar D | |||||||||
| Funding support |  Canada, 2 items Canada, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2021 Journal: Cell Rep / Year: 2021Title: Structural analysis of receptor binding domain mutations in SARS-CoV-2 variants of concern that modulate ACE2 and antibody binding. Authors: Dhiraj Mannar / James W Saville / Xing Zhu / Shanti S Srivastava / Alison M Berezuk / Steven Zhou / Katharine S Tuttle / Andrew Kim / Wei Li / Dimiter S Dimitrov / Sriram Subramaniam /   Abstract: The recently emerged severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) Beta (B.1.351) and Gamma (P.1) variants of concern (VoCs) include a key mutation (N501Y) found in the Alpha (B.1.1.7) ...The recently emerged severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) Beta (B.1.351) and Gamma (P.1) variants of concern (VoCs) include a key mutation (N501Y) found in the Alpha (B.1.1.7) variant that enhances affinity of the spike protein for its receptor, angiotensin-converting enzyme 2 (ACE2). Additional mutations are found in these variants at residues 417 and 484 that appear to promote antibody evasion. In contrast, the Epsilon variants (B.1.427/429) lack the N501Y mutation yet exhibit antibody evasion. We have engineered spike proteins to express these receptor binding domain (RBD) VoC mutations either in isolation or in different combinations and analyze the effects using biochemical assays and cryoelectron microscopy (cryo-EM) structural analyses. Overall, our findings suggest that the emergence of new SARS-CoV-2 variant spikes can be rationalized as the result of mutations that confer increased ACE2 affinity, increased antibody evasion, or both, providing a framework to dissect the molecular factors that drive VoC evolution. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25513.map.gz emd_25513.map.gz | 122.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25513-v30.xml emd-25513-v30.xml emd-25513.xml emd-25513.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25513_fsc.xml emd_25513_fsc.xml | 13.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_25513.png emd_25513.png | 94.7 KB | ||
| Filedesc metadata |  emd-25513.cif.gz emd-25513.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25513 http://ftp.pdbj.org/pub/emdb/structures/EMD-25513 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25513 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25513 | HTTPS FTP |
-Related structure data
| Related structure data |  7sy1MC  7sxrC  7sxsC  7sxtC  7sxuC  7sxvC  7sxwC  7sxxC  7sxyC  7sxzC  7sy0C  7sy2C  7sy3C  7sy4C  7sy5C  7sy6C  7sy7C  7sy8C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25513.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25513.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Global refinement map of SARS-CoV-2 D614G,N501Y mutant spike protein ectodomain bound to human ACE2 ectodomain | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SARS-CoV-2 D614G,N501Y mutant spike protein ectodomain bound to h...
| Entire | Name: SARS-CoV-2 D614G,N501Y mutant spike protein ectodomain bound to human ACE2 ectodomain |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 D614G,N501Y mutant spike protein ectodomain bound to h...
| Supramolecule | Name: SARS-CoV-2 D614G,N501Y mutant spike protein ectodomain bound to human ACE2 ectodomain type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 142.418469 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF ...String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF EYVSQPFLMD LEGKQGNFKN LREFVFKNID GYFKIYSKHT PINLVRDLPQ GFSALEPLVD LPIGINITRF QT LLALHRS YLTPGDSSSG WTAGAAAYYV GYLQPRTFLL KYNENGTITD AVDCALDPLS ETKCTLKSFT VEKGIYQTSN FRV QPTESI VRFPNITNLC PFGEVFNATR FASVYAWNRK RISNCVADYS VLYNSASFST FKCYGVSPTK LNDLCFTNVY ADSF VIRGD EVRQIAPGQT GKIADYNYKL PDDFTGCVIA WNSNNLDSKV GGNYNYLYRL FRKSNLKPFE RDISTEIYQA GSTPC NGVE GFNCYFPLQS YGFQPTYGVG YQPYRVVVLS FELLHAPATV CGPKKSTNLV KNKCVNFNFN GLTGTGVLTE SNKKFL PFQ QFGRDIADTT DAVRDPQTLE ILDITPCSFG GVSVITPGTN TSNQVAVLYQ GVNCTEVPVA IHADQLTPTW RVYSTGS NV FQTRAGCLIG AEHVNNSYEC DIPIGAGICA SYQTQTNSPG SASSVASQSI IAYTMSLGAE NSVAYSNNSI AIPTNFTI S VTTEILPVSM TKTSVDCTMY ICGDSTECSN LLLQYGSFCT QLNRALTGIA VEQDKNTQEV FAQVKQIYKT PPIKDFGGF NFSQILPDPS KPSKRSPIED LLFNKVTLAD AGFIKQYGDC LGDIAARDLI CAQKFNGLTV LPPLLTDEMI AQYTSALLAG TITSGWTFG AGPALQIPFP MQMAYRFNGI GVTQNVLYEN QKLIANQFNS AIGKIQDSLS STPSALGKLQ DVVNQNAQAL N TLVKQLSS NFGAISSVLN DILSRLDPPE AEVQIDRLIT GRLQSLQTYV TQQLIRAAEI RASANLAATK MSECVLGQSK RV DFCGKGY HLMSFPQSAP HGVVFLHVTY VPAQEKNFTT APAICHDGKA HFPREGVFVS NGTHWFVTQR NFYEPQIITT DNT FVSGNC DVVIGIVNNT VYDPLQPELD SFKEELDKYF KNHTSPDVDL GDISGINASV VNIQKEIDRL NEVAKNLNES LIDL QELGK YEQGSGYIPE APRDGQAYVR KDGEWVLLST FLGRSLEVLF QGPGHHHHHH HHSAWSHPQF EKGGGSGGGG SGGSA WSHP QFEK UniProtKB: Spike glycoprotein |
-Macromolecule #2: Processed angiotensin-converting enzyme 2
| Macromolecule | Name: Processed angiotensin-converting enzyme 2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 70.386992 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSTIEEQAKT FLDKFNHEAE DLFYQSSLAS WNYNTNITEE NVQNMNNAGD KWSAFLKEQS TLAQMYPLQE IQNLTVKLQL QALQQNGSS VLSEDKSKRL NTILNTMSTI YSTGKVCNPD NPQECLLLEP GLNEIMANSL DYNERLWAWE SWRSEVGKQL R PLYEEYVV ...String: QSTIEEQAKT FLDKFNHEAE DLFYQSSLAS WNYNTNITEE NVQNMNNAGD KWSAFLKEQS TLAQMYPLQE IQNLTVKLQL QALQQNGSS VLSEDKSKRL NTILNTMSTI YSTGKVCNPD NPQECLLLEP GLNEIMANSL DYNERLWAWE SWRSEVGKQL R PLYEEYVV LKNEMARANH YEDYGDYWRG DYEVNGVDGY DYSRGQLIED VEHTFEEIKP LYEHLHAYVR AKLMNAYPSY IS PIGCLPA HLLGDMWGRF WTNLYSLTVP FGQKPNIDVT DAMVDQAWDA QRIFKEAEKF FVSVGLPNMT QGFWENSMLT DPG NVQKAV CHPTAWDLGK GDFRILMCTK VTMDDFLTAH HEMGHIQYDM AYAAQPFLLR NGANEGFHEA VGEIMSLSAA TPKH LKSIG LLSPDFQEDN ETEINFLLKQ ALTIVGTLPF TYMLEKWRWM VFKGEIPKDQ WMKKWWEMKR EIVGVVEPVP HDETY CDPA SLFHVSNDYS FIRYYTRTLY QFQFQEALCQ AAKHEGPLHK CDISNSTEAG QKLFNMLRLG KSEPWTLALE NVVGAK NMN VRPLLNYFEP LFTWLKDQNK NSFVGWSTDW SPYADHHHHH HHH UniProtKB: Angiotensin-converting enzyme 2 |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 28 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)