[English] 日本語
 Yorodumi
Yorodumi- EMDB-25472: Cryo-EM structure of Arabidopsis Ago10-guide-target RNA complex i... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Arabidopsis Ago10-guide-target RNA complex in a central duplex conformation | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | miRNA / siRNA / RNA silencing / Argonaute / plant / GENE REGULATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of shoot apical meristem development / regulation of meristem structural organization / primary shoot apical meristem specification / miRNA metabolic process / regulatory ncRNA-mediated gene silencing / miRNA binding / plastid / somatic stem cell population maintenance / RNA endonuclease activity / regulation of translation ...regulation of shoot apical meristem development / regulation of meristem structural organization / primary shoot apical meristem specification / miRNA metabolic process / regulatory ncRNA-mediated gene silencing / miRNA binding / plastid / somatic stem cell population maintenance / RNA endonuclease activity / regulation of translation / defense response to virus / ribonucleoprotein complex / RNA binding / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
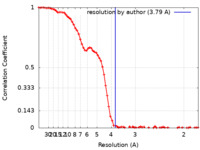
| Method | single particle reconstruction / cryo EM / Resolution: 3.79 Å | |||||||||
 Authors Authors | Xiao Y / MacRae IJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2022 Journal: Nucleic Acids Res / Year: 2022Title: The molecular mechanism of microRNA duplex selectivity of Arabidopsis ARGONAUTE10. Authors: Yao Xiao / Ian J MacRae /  Abstract: Small RNAs (sRNAs), including microRNAs (miRNAs) and small interfering RNAs (siRNAs), are essential gene regulators for plant and animal development. The loading of sRNA duplexes into the proper ...Small RNAs (sRNAs), including microRNAs (miRNAs) and small interfering RNAs (siRNAs), are essential gene regulators for plant and animal development. The loading of sRNA duplexes into the proper ARGONAUTE (AGO) protein is a key step to forming a functional silencing complex. In Arabidopsis thaliana, the specific loading of miR166/165 into AGO10 (AtAGO10) is critical for the maintenance of the shoot apical meristem, the source of all shoot organs, but the mechanism by which AtAGO10 distinguishes miR166/165 from other cellular miRNAs is not known. Here, we show purified AtAGO10 alone lacks loading selectivity towards miR166/165 duplexes. However, phosphate and HSP chaperone systems reshape the selectivity of AtAGO10 to its physiological substrates. A loop in the AtAGO10 central cleft is essential for recognizing specific mismatches opposite the guide strand 3' region in miR166/165 duplexes. Replacing this loop with the equivalent loop from Homo sapiens AGO2 (HsAGO2) changes AtAGO10 miRNA loading behavior such that 3' region mismatches are ignored and mismatches opposite the guide 5' end instead drive loading, as in HsAGO2. Thus, this study uncovers the molecular mechanism underlying the miR166/165 selectivity of AtAGO10, essential for plant development, and provides new insights into how miRNA duplex structures are recognized for sRNA sorting. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25472.map.gz emd_25472.map.gz | 23.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25472-v30.xml emd-25472-v30.xml emd-25472.xml emd-25472.xml | 24 KB 24 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25472_fsc.xml emd_25472_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_25472.png emd_25472.png | 64.9 KB | ||
| Filedesc metadata |  emd-25472.cif.gz emd-25472.cif.gz | 7.3 KB | ||
| Others |  emd_25472_additional_1.map.gz emd_25472_additional_1.map.gz emd_25472_half_map_1.map.gz emd_25472_half_map_1.map.gz emd_25472_half_map_2.map.gz emd_25472_half_map_2.map.gz | 20.8 MB 23.4 MB 23.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25472 http://ftp.pdbj.org/pub/emdb/structures/EMD-25472 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25472 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25472 | HTTPS FTP |
-Related structure data
| Related structure data |  7swfMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25472.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25472.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.91 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_25472_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
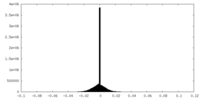
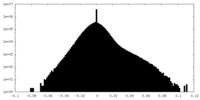
| Density Histograms |
-Half map: #2
| File | emd_25472_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_25472_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : AtAgo10-guide-target RNA complex
| Entire | Name: AtAgo10-guide-target RNA complex |
|---|---|
| Components |
|
-Supramolecule #1: AtAgo10-guide-target RNA complex
| Supramolecule | Name: AtAgo10-guide-target RNA complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: The complex of Arabidopsis Argonaute10 with a synthetic guide RNA and a target pairing to 2-16nt of guide guide |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Protein argonaute 10
| Macromolecule | Name: Protein argonaute 10 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 110.981633 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPIRQMKDSS ETHLVIKTQP LKHHNPKTVQ NGKIPPPSPS PVTVTTPATV TQSQASSPSP PSKNRSRRRN RGGRKSDQGD VCMRPSSRP RKPPPPSQTT SSAVSVATAG EIVAVNHQMQ MGVRKNSNFA PRPGFGTLGT KCIVKANHFL ADLPTKDLNQ Y DVTITPEV ...String: MPIRQMKDSS ETHLVIKTQP LKHHNPKTVQ NGKIPPPSPS PVTVTTPATV TQSQASSPSP PSKNRSRRRN RGGRKSDQGD VCMRPSSRP RKPPPPSQTT SSAVSVATAG EIVAVNHQMQ MGVRKNSNFA PRPGFGTLGT KCIVKANHFL ADLPTKDLNQ Y DVTITPEV SSKSVNRAII AELVRLYKES DLGRRLPAYD GRKSLYTAGE LPFTWKEFSV KIVDEDDGII NGPKRERSYK VA IKFVARA NMHHLGEFLA GKRADCPQEA VQILDIVLRE LSVKRFCPVG RSFFSPDIKT PQRLGEGLES WCGFYQSIRP TQM GLSLNI DMASAAFIEP LPVIEFVAQL LGKDVLSKPL SDSDRVKIKK GLRGVKVEVT HRANVRRKYR VAGLTTQPTR ELMF PVDEN CTMKSVIEYF QEMYGFTIQH THLPCLQVGN QKKASYLPME ACKIVEGQRY TKRLNEKQIT ALLKVTCQRP RDREN DILR TVQHNAYDQD PYAKEFGMNI SEKLASVEAR ILPAPWLKYH ENGKEKDCLP QVGQWNMMNK KMINGMTVSR WACVNF SRS VQENVARGFC NELGQMCEVS GMEFNPEPVI PIYSARPDQV EKALKHVYHT SMNKTKGKEL ELLLAILPDN NGSLYGD LK RICETELGLI SQCCLTKHVF KISKQYLANV SLKINVKMGG RNTVLVDAIS CRIPLVSDIP TIIFGADVTH PENGEESS P SIAAVVASQD WPEVTKYAGL VCAQAHRQEL IQDLYKTWQD PVRGTVSGGM IRDLLISFRK ATGQKPLRII FYRAGVSEG QFYQVLLYEL DAIRKACASL EPNYQPPVTF IVVQKRHHTR LFANNHRDKN STDRSGNILP GTVVDTKICH PTEFDFYLCS HAGIQGTSR PAHYHVLWDE NNFTADGIQS LTNNLCYTYA RCTRSVSIVP PAYYAHLAAF RARFYLEPEI MQDNGSPGKK N TKTTTVGD VGVKPLPALK ENVKRVMFYC UniProtKB: Protein argonaute 10 |
-Macromolecule #2: RNA (5'-R(P*UP*GP*GP*AP*GP*UP*GP*UP*GP*AP*CP*AP*AP*UP*GP*GP*U)-3')
| Macromolecule | Name: RNA (5'-R(P*UP*GP*GP*AP*GP*UP*GP*UP*GP*AP*CP*AP*AP*UP*GP*GP*U)-3') type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 6.788021 KDa |
| Sequence | String: UGGAGUGUGA CAAUGGUGUU U |
-Macromolecule #3: RNA (5'-R(P*CP*CP*AP*UP*UP*GP*UP*CP*AP*CP*AP*CP*UP*CP*CP*AP*A)-3')
| Macromolecule | Name: RNA (5'-R(P*CP*CP*AP*UP*UP*GP*UP*CP*AP*CP*AP*CP*UP*CP*CP*AP*A)-3') type: rna / ID: 3 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 5.636419 KDa |
| Sequence | String: CCAUUGUCAC ACUCCAAA |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 2049 / Average electron dose: 66.88 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.7 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 45000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)