+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
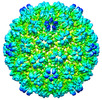
| Title | Procapsid of bacteriophage lambda | |||||||||
 Map data Map data | Low resolution phage lambda procapsid map. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Escherichia virus Lambda Escherichia virus Lambda | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.0 Å | |||||||||
 Authors Authors | Maruthi K / Prokhorov NS / Morais MC | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Assembly Incompetent Major Capsid Protein Reveals Elementary and Transient Interactions with Scaffolding Chaparone that Nucleate Viral Shell Assembly Authors: Davis CR / Churchill ME / Backos D / Maruthi K / Prokhorov NS / Morais MC / Catalano CE | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25182.map.gz emd_25182.map.gz | 21.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25182-v30.xml emd-25182-v30.xml emd-25182.xml emd-25182.xml | 7.9 KB 7.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25182.png emd_25182.png | 265.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25182 http://ftp.pdbj.org/pub/emdb/structures/EMD-25182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25182 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25182.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25182.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Low resolution phage lambda procapsid map. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.8 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Procapsid of bacteriophage lambda
| Entire | Name: Procapsid of bacteriophage lambda |
|---|---|
| Components |
|
-Supramolecule #1: Procapsid of bacteriophage lambda
| Supramolecule | Name: Procapsid of bacteriophage lambda / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Escherichia virus Lambda Escherichia virus Lambda |
-Macromolecule #1: Major Capsid Protein
| Macromolecule | Name: Major Capsid Protein / type: protein_or_peptide / ID: 1 / Enantiomer: DEXTRO |
|---|---|
| Sequence | String: MSMYTTAQLL AANEQKFKFD PLFLRLFFRE SYPFTTEKVY LSQIPGLVNM ALYVSPIVSG EVIRSRGGS TSEFTPGYVK PKHEVNPQMT LRRLPDEDPQ NLADPAYRRR RIIMQNMRDE E LAIAQVEE MQAVSAVLKG KYTMTGEAFD PVEVDMGRSE ENNITQSGGT ...String: MSMYTTAQLL AANEQKFKFD PLFLRLFFRE SYPFTTEKVY LSQIPGLVNM ALYVSPIVSG EVIRSRGGS TSEFTPGYVK PKHEVNPQMT LRRLPDEDPQ NLADPAYRRR RIIMQNMRDE E LAIAQVEE MQAVSAVLKG KYTMTGEAFD PVEVDMGRSE ENNITQSGGT EWSKRDKSTY DP TDDIEAY ALNASGVVNI IVFDPKGWAL FRSFKAVKEK LDTRRGSNSE LETAVKDLGK AVS YKGMYG DVAIVVYSGQ YVENGVKKNF LPDNTMVLGN TQARGLRTYG CIQDADAQRE GINA SARYP KNWVTTGDPA REFTMIQSAP LMLLADPDEF VSVQLA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2200FS |
|---|---|
| Image recording | Film or detector model: DIRECT ELECTRON DE-20 (5k x 3k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 8.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 1253 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




















